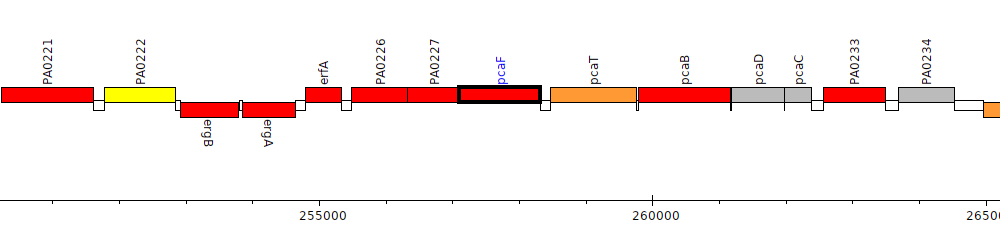
Pseudomonas aeruginosa PAO1, PA0228 (pcaF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044248 | cellular catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0015976 | carbon utilization | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044255 | cellular lipid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000429
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019619 | 3,4-dihydroxybenzoate catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02430
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016746 | transferase activity, transferring acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53901
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Aromatic compound catabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00362 | Benzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.40.47.10:FF:000010 | Acetyl-CoA acetyltransferase (Thiolase) | - | - | 1 | 400 | 0.0 |
| PANTHER | PTHR18919 | ACETYL-COA C-ACYLTRANSFERASE | - | - | 1 | 401 | 5.9E-129 |
| Pfam | PF00108 | Thiolase, N-terminal domain | IPR020616 | Thiolase, N-terminal | 5 | 268 | 4.4E-79 |
| NCBIfam | TIGR02430 | JCVI: 3-oxoadipyl-CoA thiolase | IPR012793 | Beta-ketoadipyl CoA thiolase | 3 | 401 | 0.0 |
| SUPERFAMILY | SSF53901 | Thiolase-like | IPR016039 | Thiolase-like | 277 | 400 | 3.12E-45 |
| Pfam | PF02803 | Thiolase, C-terminal domain | IPR020617 | Thiolase, C-terminal | 277 | 400 | 7.4E-43 |
| NCBIfam | TIGR01930 | JCVI: acetyl-CoA C-acyltransferase | IPR002155 | Thiolase | 7 | 399 | 0.0 |
| CDD | cd00751 | thiolase | IPR002155 | Thiolase | 6 | 400 | 0.0 |
| Gene3D | G3DSA:3.40.47.10 | - | IPR016039 | Thiolase-like | 1 | 401 | 0.0 |
| PIRSF | PIRSF000429 | Ac-CoA_Ac_transf | IPR002155 | Thiolase | 1 | 401 | 0.0 |
| SUPERFAMILY | SSF53901 | Thiolase-like | IPR016039 | Thiolase-like | 4 | 276 | 4.51E-88 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.