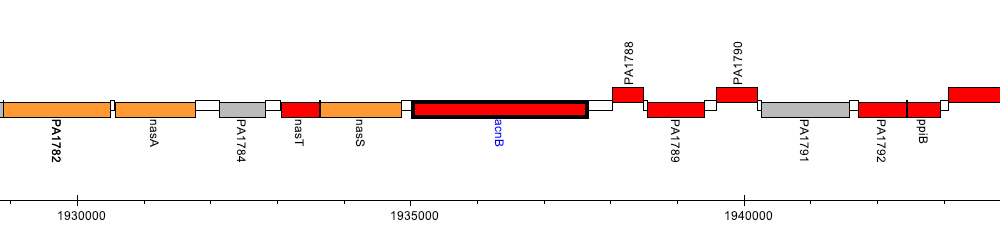
Pseudomonas aeruginosa PAO1, PA1787 (acnB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0003994 | aconitate hydratase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8000525 | Reviewed by curator |
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:PF11791
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005829 | cytosol |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF036687
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003994 | aconitate hydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF11791
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF036687
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00640 | Propanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | GLYCOLYSIS-TCA-GLYOX-BYPASS | superpathway of glycolysis, pyruvate dehydrogenase, TCA, and glyoxylate bypass | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Reductive carboxylate cycle (CO2 fixation) |
ECO:0000037
not_recorded |
|||
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glyoxylate and dicarboxylate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | ANARESP1-PWY | anaerobic respiration | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P22-PWY | acetyl-CoA assimilation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | TCA | TCA cycle I (prokaryotic) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00020 | Citrate cycle (TCA cycle) | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | GLYOXYLATE-BYPASS | glyoxylate cycle | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | TCA-GLYOX-BYPASS | superpathway of glyoxylate bypass and TCA | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P106-PWY | serine-isocitrate lyase pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01210 | 2-Oxocarboxylic acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Citrate cycle (TCA cycle) |
ECO:0000037
not_recorded |
|||
| PseudoCyc | FERMENTATION-PWY | mixed acid fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00117 | JCVI: bifunctional aconitate hydratase 2/2-methylisocitrate dehydratase | IPR004406 | Aconitase B | 1 | 857 | 0.0 |
| FunFam | G3DSA:3.20.19.10:FF:000004 | Aconitate hydratase B | - | - | 162 | 361 | 1.2E-115 |
| Gene3D | G3DSA:1.25.40.310 | - | IPR036288 | Aconitase B, HEAT-like domain superfamily | 1 | 161 | 1.7E-75 |
| Gene3D | G3DSA:3.20.19.10 | Aconitase, domain 4 | IPR015928 | Aconitase/3-isopropylmalate dehydratase, swivel | 162 | 361 | 1.8E-97 |
| PANTHER | PTHR43160 | ACONITATE HYDRATASE B | - | - | 245 | 824 | 0.0 |
| FunFam | G3DSA:1.25.40.310:FF:000001 | Aconitate hydratase B | - | - | 1 | 161 | 9.9E-84 |
| SUPERFAMILY | SSF53732 | Aconitase iron-sulfur domain | IPR036008 | Aconitase, iron-sulfur domain | 379 | 858 | 0.0 |
| Gene3D | G3DSA:3.30.499.10 | Aconitase, domain 3 | IPR015931 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha, subdomain 1/3 | 686 | 867 | 4.4E-66 |
| SUPERFAMILY | SSF74778 | Aconitase B, N-terminal domain | IPR036288 | Aconitase B, HEAT-like domain superfamily | 1 | 160 | 4.05E-69 |
| FunFam | G3DSA:3.30.499.10:FF:000008 | Aconitate hydratase B | - | - | 686 | 868 | 5.1E-101 |
| Pfam | PF06434 | Aconitate hydratase 2 N-terminus | IPR015929 | Aconitase B, swivel | 168 | 382 | 2.8E-94 |
| PIRSF | PIRSF036687 | AcnB | IPR004406 | Aconitase B | 1 | 862 | 0.0 |
| Pfam | PF00330 | Aconitase family (aconitate hydratase) | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 475 | 821 | 2.1E-41 |
| CDD | cd01576 | AcnB_Swivel | IPR015929 | Aconitase B, swivel | 171 | 311 | 1.30507E-73 |
| Pfam | PF11791 | Aconitate B N-terminal domain | IPR015933 | Aconitase B, HEAT-like domain | 4 | 156 | 6.1E-67 |
| FunFam | G3DSA:3.30.499.10:FF:000001 | Aconitate hydratase B | - | - | 405 | 532 | 1.3E-84 |
| CDD | cd01581 | AcnB | - | - | 384 | 823 | 0.0 |
| Gene3D | G3DSA:3.40.1060.10 | Aconitase, Domain 2 | IPR015932 | Aconitase, domain 2 | 382 | 684 | 0.0 |
| Gene3D | G3DSA:3.30.499.10 | Aconitase, domain 3 | IPR015931 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha, subdomain 1/3 | 405 | 532 | 0.0 |
| SUPERFAMILY | SSF52016 | LeuD/IlvD-like | - | - | 161 | 372 | 3.57E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.