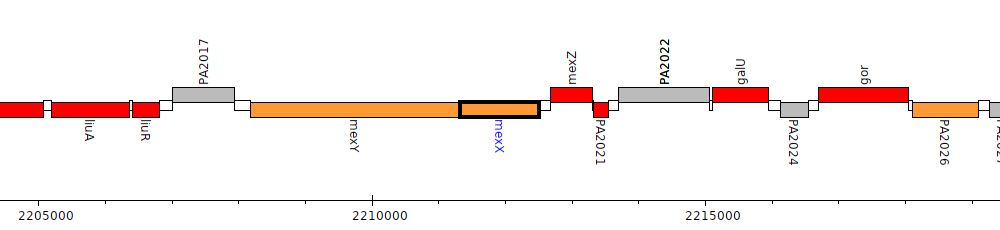
Pseudomonas aeruginosa PAO1, PA2019 (mexX)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
10952562 | Reviewed by curator |
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Experiment | ECO:0000269 experimental evidence used in manual assertion |
10952562 | Reviewed by curator |
| Molecular Function | GO:0042910 | xenobiotic transmembrane transporter activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9925549 | Reviewed by curator |
| Molecular Function | GO:0042910 | xenobiotic transmembrane transporter activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
10543738 | Reviewed by curator |
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Experiment | ECO:0000269 experimental evidence used in manual assertion |
16030228 | Reviewed by curator |
| Biological Process | GO:0046677 | response to antibiotic | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0042493 | response to drug | Inferred from Experiment | ECO:0000269 experimental evidence used in manual assertion |
16030228 | Reviewed by curator |
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF00529
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01501 | beta-Lactam resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.40.30.170 | - | - | - | 58 | 285 | 1.8E-68 |
| NCBIfam | TIGR01730 | JCVI: efflux RND transporter periplasmic adaptor subunit | IPR006143 | RND efflux pump, membrane fusion protein | 42 | 375 | 1.5E-70 |
| Gene3D | G3DSA:1.10.287.470 | Helix hairpin bin | - | - | 99 | 172 | 1.8E-68 |
| FunFam | G3DSA:2.40.420.20:FF:000040 | Multidrug efflux RND transporter periplasmic adaptor subunit MexX | - | - | 286 | 376 | 8.7E-60 |
| Gene3D | G3DSA:2.40.50.100 | - | - | - | 64 | 208 | 1.8E-68 |
| Coils | Coil | Coil | - | - | 143 | 170 | - |
| PANTHER | PTHR30158 | ACRA/E-RELATED COMPONENT OF DRUG EFFLUX TRANSPORTER | - | - | 2 | 387 | 2.1E-117 |
| Pfam | PF00529 | Cation efflux system protein CusB domain 1 | IPR043602 | Cation efflux system protein CusB, domain 1 | 40 | 360 | 4.7E-12 |
| Pfam | PF16576 | Barrel-sandwich domain of CusB or HlyD membrane-fusion | IPR032317 | RND efflux pump, membrane fusion protein, barrel-sandwich domain | 62 | 294 | 1.4E-10 |
| Gene3D | G3DSA:2.40.420.20 | - | - | - | 286 | 376 | 2.2E-15 |
| SUPERFAMILY | SSF111369 | HlyD-like secretion proteins | - | - | 53 | 300 | 4.71E-72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.