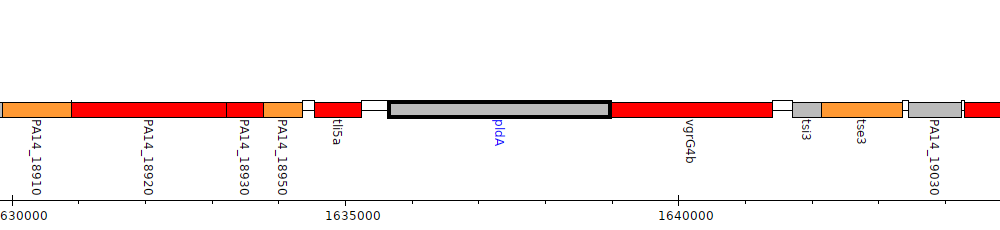
Pseudomonas aeruginosa UCBPP-PA14, PA14_18970 (pldA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00614
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00614 | Phospholipase D Active site motif | IPR001736 | Phospholipase D/Transphosphatidylase | 168 | 193 | 1.6E-7 |
| SMART | SM00155 | pld_4 | IPR001736 | Phospholipase D/Transphosphatidylase | 854 | 881 | 3.7E-6 |
| PANTHER | PTHR18896 | PHOSPHOLIPASE D | IPR015679 | Phospholipase D family | 41 | 975 | 3.8E-94 |
| SUPERFAMILY | SSF56024 | Phospholipase D/nuclease | - | - | 46 | 499 | 5.16E-25 |
| Gene3D | G3DSA:3.30.870.10 | Endonuclease Chain A | - | - | 433 | 516 | 1.3E-6 |
| Pfam | PF00614 | Phospholipase D Active site motif | IPR001736 | Phospholipase D/Transphosphatidylase | 855 | 881 | 3.4E-6 |
| Gene3D | G3DSA:3.30.870.10 | Endonuclease Chain A | - | - | 565 | 905 | 2.7E-22 |
| CDD | cd09141 | PLDc_vPLD1_2_yPLD_like_2 | - | - | 604 | 898 | 1.23989E-73 |
| Gene3D | G3DSA:3.30.870.10 | Endonuclease Chain A | - | - | 51 | 213 | 2.0E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1084 | 1103 | - |
| SMART | SM00155 | pld_4 | IPR001736 | Phospholipase D/Transphosphatidylase | 166 | 193 | 0.0016 |
| CDD | cd09104 | PLDc_vPLD1_2_like_1 | - | - | 51 | 204 | 1.80748E-29 |
| SUPERFAMILY | SSF56024 | Phospholipase D/nuclease | - | - | 610 | 929 | 8.63E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.