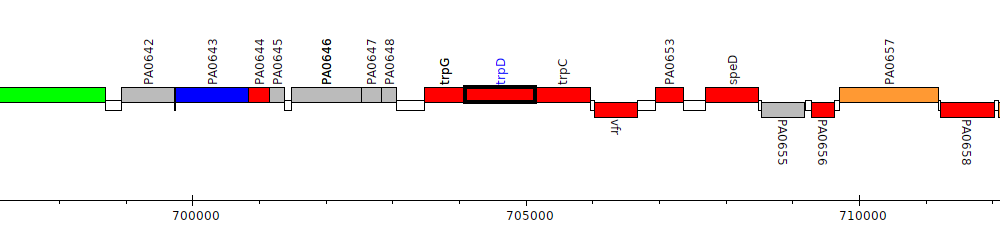
Pseudomonas aeruginosa PAO1, PA0650 (trpD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0000162 | tryptophan biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43285
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016757 | transferase activity, transferring glycosyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF00591
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004048 | anthranilate phosphoribosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43285
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | TRPSYN-PWY | L-tryptophan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | COMPLETE-ARO-PWY | superpathway of aromatic amino acid biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Phenylalanine, tyrosine and tryptophan biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | ALL-CHORISMATE-PWY | superpathway of chorismate metabolism | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00591 | Glycosyl transferase family, a/b domain | IPR000312 | Glycosyl transferase, family 3 | 75 | 329 | 1.3E-101 |
| SUPERFAMILY | SSF52418 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain | IPR035902 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain superfamily | 75 | 340 | 1.57E-87 |
| Gene3D | G3DSA:3.40.1030.10 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain | IPR035902 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain superfamily | 73 | 347 | 1.8E-103 |
| FunFam | G3DSA:3.40.1030.10:FF:000002 | Anthranilate phosphoribosyltransferase | - | - | 73 | 342 | 1.9E-103 |
| Gene3D | G3DSA:1.20.970.10 | Transferase, Pyrimidine Nucleoside Phosphorylase; Chain C | - | - | 1 | 70 | 2.5E-29 |
| NCBIfam | TIGR01245 | JCVI: anthranilate phosphoribosyltransferase | IPR005940 | Anthranilate phosphoribosyl transferase | 7 | 338 | 1.4E-120 |
| Coils | Coil | Coil | - | - | 7 | 27 | - |
| SUPERFAMILY | SSF47648 | Nucleoside phosphorylase/phosphoribosyltransferase N-terminal domain | IPR036320 | Glycosyl transferase family 3, N-terminal domain superfamily | 1 | 67 | 2.88E-21 |
| FunFam | G3DSA:1.20.970.10:FF:000006 | Anthranilate phosphoribosyltransferase | - | - | 1 | 70 | 8.2E-32 |
| Pfam | PF02885 | Glycosyl transferase family, helical bundle domain | IPR017459 | Glycosyl transferase family 3, N-terminal domain | 6 | 65 | 2.2E-19 |
| PANTHER | PTHR43285 | ANTHRANILATE PHOSPHORIBOSYLTRANSFERASE | IPR005940 | Anthranilate phosphoribosyl transferase | 2 | 341 | 3.1E-107 |
| Hamap | MF_00211 | Anthranilate phosphoribosyltransferase [trpD]. | IPR005940 | Anthranilate phosphoribosyl transferase | 5 | 339 | 37.749939 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.