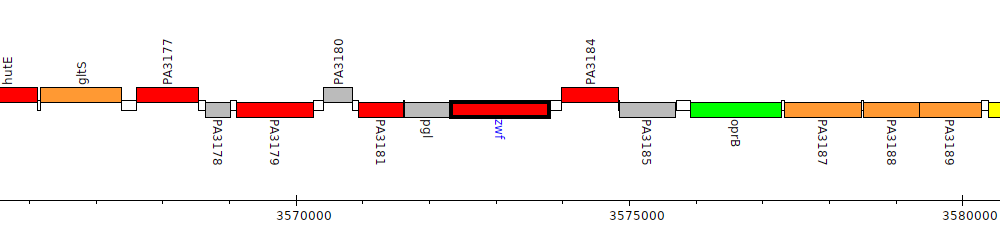
Pseudomonas aeruginosa PAO1, PA3183 (zwf)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044248 | cellular catabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0015976 | carbon utilization | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0008734 | L-aspartate oxidase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004345 | glucose-6-phosphate dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02781
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006006 | glucose metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050661 | NADP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016614 | oxidoreductase activity, acting on CH-OH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00480 | Glutathione metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P122-PWY | heterolactic fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | OXIDATIVEPENT-PWY | pentose phosphate pathway (oxidative branch) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PENTOSE-P-PWY | pentose phosphate pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Pentose phosphate cycle |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Glutathione metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | GLYCOLYSIS-E-D | superpathway of glycolysis and Entner-Doudoroff | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00079 | Glucose-6-phosphate dehydrogenase signature | IPR001282 | Glucose-6-phosphate dehydrogenase | 325 | 351 | 1.7E-49 |
| SUPERFAMILY | SSF55347 | Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domain | - | - | 179 | 486 | 4.58E-114 |
| PRINTS | PR00079 | Glucose-6-phosphate dehydrogenase signature | IPR001282 | Glucose-6-phosphate dehydrogenase | 164 | 192 | 1.7E-49 |
| NCBIfam | TIGR00871 | JCVI: glucose-6-phosphate dehydrogenase | IPR001282 | Glucose-6-phosphate dehydrogenase | 8 | 485 | 0.0 |
| Pfam | PF02781 | Glucose-6-phosphate dehydrogenase, C-terminal domain | IPR022675 | Glucose-6-phosphate dehydrogenase, C-terminal | 187 | 486 | 9.4E-120 |
| PANTHER | PTHR23429 | GLUCOSE-6-PHOSPHATE 1-DEHYDROGENASE G6PD | IPR001282 | Glucose-6-phosphate dehydrogenase | 9 | 484 | 0.0 |
| PRINTS | PR00079 | Glucose-6-phosphate dehydrogenase signature | IPR001282 | Glucose-6-phosphate dehydrogenase | 216 | 233 | 1.7E-49 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 7 | 187 | 2.9E-53 |
| PRINTS | PR00079 | Glucose-6-phosphate dehydrogenase signature | IPR001282 | Glucose-6-phosphate dehydrogenase | 140 | 153 | 1.7E-49 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 5 | 172 | 6.0E-41 |
| Pfam | PF00479 | Glucose-6-phosphate dehydrogenase, NAD binding domain | IPR022674 | Glucose-6-phosphate dehydrogenase, NAD-binding | 13 | 184 | 4.3E-51 |
| PRINTS | PR00079 | Glucose-6-phosphate dehydrogenase signature | IPR001282 | Glucose-6-phosphate dehydrogenase | 234 | 250 | 1.7E-49 |
| Hamap | MF_00966 | Glucose-6-phosphate 1-dehydrogenase [zwf]. | IPR001282 | Glucose-6-phosphate dehydrogenase | 8 | 487 | 89.891327 |
| PIRSF | PIRSF000110 | G6PD | IPR001282 | Glucose-6-phosphate dehydrogenase | 2 | 488 | 0.0 |
| FunFam | G3DSA:3.30.360.10:FF:000011 | Glucose-6-phosphate 1-dehydrogenase | - | - | 174 | 477 | 1.8E-119 |
| Gene3D | G3DSA:3.30.360.10 | Dihydrodipicolinate Reductase; domain 2 | - | - | 173 | 487 | 1.3E-122 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.