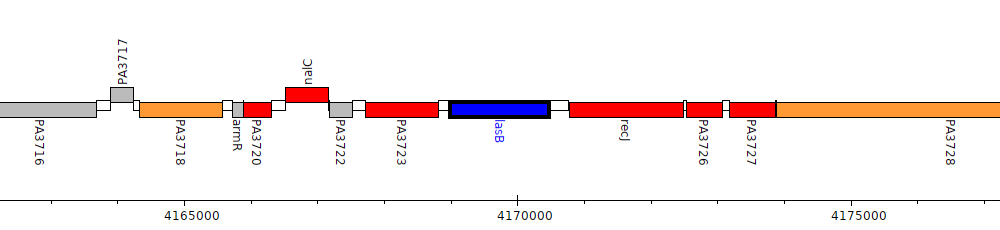
Pseudomonas aeruginosa PAO1, PA3724 (lasB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0071978 | bacterial-type flagellum-dependent swarming motility | Inferred from Mutant Phenotype | ECO:0000016 loss-of-function mutant phenotype evidence |
18245294 | Reviewed by curator |
| Biological Process | GO:0060309 | elastin catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2842313 | Reviewed by curator |
| Biological Process | GO:0043952 | protein transport by the Sec complex | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9642203 | Reviewed by curator |
| Biological Process | GO:0015628 | protein secretion by the type II secretion system | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
9642203 | Reviewed by curator |
| Biological Process | GO:0052051 | obsolete interaction with host via protein secreted by type II secretion system | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
15123664 | Reviewed by curator |
| Biological Process | GO:0044010 | single-species biofilm formation | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18245294 | Reviewed by curator |
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0044010 | single-species biofilm formation | Inferred from Mutant Phenotype | ECO:0000016 loss-of-function mutant phenotype evidence |
18245294 | Reviewed by curator |
| Biological Process | GO:0006508 | proteolysis | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
1597429 | Reviewed by curator |
| Biological Process | GO:0060309 | elastin catabolic process | Inferred from Genetic Interaction | ECO:0000316 genetic interaction evidence used in manual assertion |
1902216 | Reviewed by curator |
| Biological Process | GO:0051542 | elastin biosynthetic process | Inferred from Genetic Interaction | ECO:0000316 genetic interaction evidence used in manual assertion |
1902216 | Reviewed by curator |
| Biological Process | GO:0071978 | bacterial-type flagellum-dependent swarming motility | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18245294 | Reviewed by curator |
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae02024 | Quorum sensing | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Xcp type II secretion system |
ECO:0000037
not_recorded |
|||
| KEGG | pae01503 | Cationic antimicrobial peptide (CAMP) resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55486 | Metalloproteases ("zincins"), catalytic domain | - | - | 199 | 494 | 1.41E-88 |
| Gene3D | G3DSA:3.10.170.10 | - | - | - | 198 | 349 | 1.8E-56 |
| Pfam | PF03413 | Peptidase propeptide and YPEB domain | IPR025711 | PepSY domain | 123 | 192 | 8.8E-9 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 336 | 352 | 5.8E-26 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 419 | 435 | 5.8E-26 |
| Pfam | PF07504 | Fungalysin/Thermolysin Propeptide Motif | IPR011096 | FTP domain | 56 | 90 | 3.0E-6 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 357 | 368 | 5.8E-26 |
| FunFam | G3DSA:3.10.170.10:FF:000002 | Elastase | - | - | 198 | 349 | 9.9E-98 |
| Gene3D | G3DSA:3.10.450.40 | - | - | - | 118 | 197 | 4.2E-22 |
| Gene3D | G3DSA:3.10.450.490 | - | - | - | 30 | 115 | 1.1E-11 |
| Gene3D | G3DSA:1.10.390.10 | Neutral Protease Domain 2 | IPR027268 | Peptidase M4/M1, CTD superfamily | 350 | 497 | 3.5E-49 |
| Pfam | PF01447 | Thermolysin metallopeptidase, catalytic domain | IPR013856 | Peptidase M4 domain | 208 | 345 | 3.4E-33 |
| CDD | cd09597 | M4_TLP | - | - | 234 | 492 | 8.53745E-101 |
| Pfam | PF02868 | Thermolysin metallopeptidase, alpha-helical domain | IPR001570 | Peptidase M4, C-terminal | 348 | 492 | 1.0E-36 |
| PRINTS | PR00730 | Thermolysin metalloprotease (M4) family signature | IPR023612 | Peptidase M4 | 302 | 322 | 5.8E-26 |
| PANTHER | PTHR33794 | BACILLOLYSIN | - | - | 34 | 492 | 7.3E-104 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.