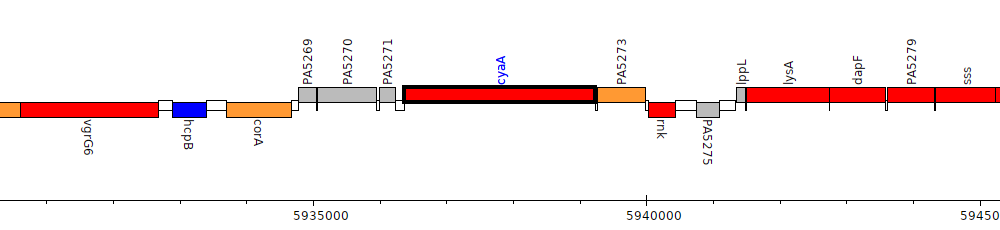
Pseudomonas aeruginosa PAO1, PA5272 (cyaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004016 | adenylate cyclase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
12586068 | Reviewed by curator |
| Biological Process | GO:0009405 | pathogenesis | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
14977975 | Reviewed by curator |
| Biological Process | GO:0006171 | cAMP biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
12586068 | Reviewed by curator |
| Molecular Function | GO:0004016 | adenylate cyclase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01295
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006171 | cAMP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01295
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF001444 | Adenylate_cycl | IPR000274 | Adenylate cyclase, class-I | 16 | 931 | 0.0 |
| PANTHER | PTHR38760 | ADENYLATE CYCLASE | IPR000274 | Adenylate cyclase, class-I | 19 | 861 | 0.0 |
| Pfam | PF01295 | Adenylate cyclase, class-I | IPR000274 | Adenylate cyclase, class-I | 244 | 844 | 6.9E-121 |
| Pfam | PF12633 | Adenylate cyclase NT domain | IPR024685 | Adenylate cyclase class-I, N-terminal domain | 21 | 217 | 2.7E-72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.