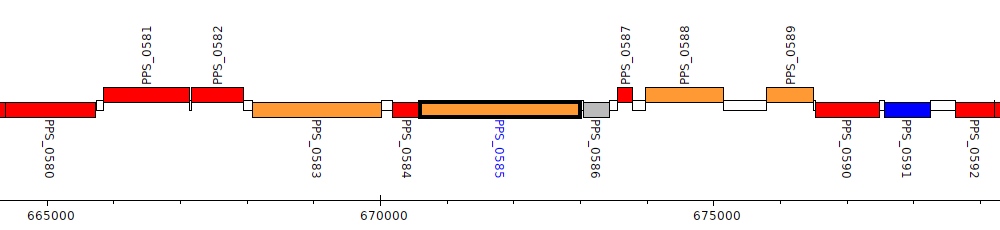
Pseudomonas putida S16, PPS_0585
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005215 | transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00003
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 481 | 698 | 3.4E-42 |
| FunFam | G3DSA:3.30.70.100:FF:000005 | Copper-exporting P-type ATPase A | - | - | 71 | 138 | 5.2E-21 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 254 | 273 | 5.4E-14 |
| CDD | cd02094 | P-type_ATPase_Cu-like | - | - | 168 | 787 | 0.0 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 74 | 136 | 9.64474E-19 |
| SUPERFAMILY | SSF81665 | Calcium ATPase, transmembrane domain M | IPR023298 | P-type ATPase, transmembrane domain superfamily | 254 | 760 | 3.53E-15 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 204 | 223 | 5.4E-14 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 70 | 138 | 4.32E-19 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 707 | 719 | 1.1E-21 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 467 | 482 | 5.4E-14 |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | - | - | 466 | 736 | 0.0 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 407 | 421 | 5.4E-14 |
| SUPERFAMILY | SSF81653 | Calcium ATPase, transduction domain A | IPR008250 | P-type ATPase, A domain superfamily | 287 | 384 | 6.93E-24 |
| NCBIfam | TIGR01525 | JCVI: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 219 | 785 | 0.0 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 75 | 133 | 1.0E-12 |
| FunFam | G3DSA:3.30.70.100:FF:000005 | Copper-exporting P-type ATPase A | - | - | 3 | 70 | 1.2E-18 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 274 | 292 | 5.4E-14 |
| NCBIfam | TIGR01511 | JCVI: copper-translocating P-type ATPase | - | - | 201 | 787 | 0.0 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 483 | 784 | 4.54E-56 |
| SFLD | SFLDF00027 | p-type atpase | IPR044492 | P-type ATPase, haloacid dehalogenase domain | 466 | 736 | 0.0 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 451 | 741 | 6.1E-58 |
| PANTHER | PTHR43520 | ATP7, ISOFORM B | - | - | 4 | 790 | 0.0 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 632 | 642 | 1.1E-21 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 71 | 137 | 4.1E-21 |
| Gene3D | G3DSA:2.70.150.10 | - | - | - | 272 | 388 | 2.1E-31 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 684 | 703 | 1.1E-21 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 10 | 66 | 1.1E-12 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 334 | 348 | 1.1E-21 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 257 | 767 | 8.8E-86 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 5 | 68 | 2.88E-17 |
| Pfam | PF00122 | E1-E2 ATPase | - | - | 285 | 464 | 1.8E-52 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 6 | 70 | 2.1E-18 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 87 | 101 | 5.4E-14 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 662 | 679 | 5.4E-14 |
| FunFam | G3DSA:2.70.150.10:FF:000020 | Copper-exporting P-type ATPase A | - | - | 272 | 388 | 1.6E-36 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 484 | 498 | 1.1E-21 |
| NCBIfam | TIGR00003 | JCVI: copper ion binding protein | IPR006122 | Heavy metal-associated domain, copper ion-binding | 8 | 68 | 9.5E-8 |
| NCBIfam | TIGR00003 | JCVI: copper ion binding protein | IPR006122 | Heavy metal-associated domain, copper ion-binding | 73 | 135 | 4.0E-11 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 8 | 68 | 4.80277E-16 |
| Gene3D | G3DSA:3.40.1110.10 | - | IPR023299 | P-type ATPase, cytoplasmic domain N | 495 | 616 | 9.7E-27 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.