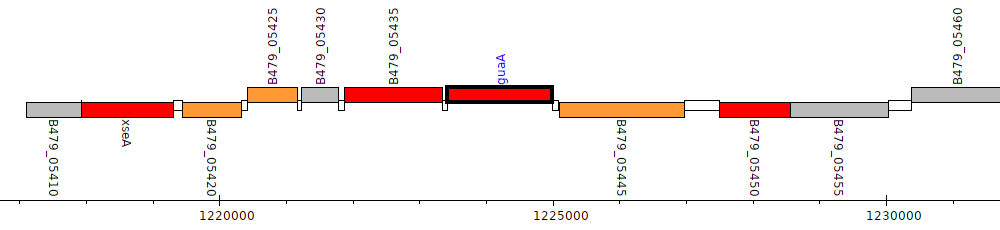
Pseudomonas putida HB3267, B479_05440 (guaA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006164 | purine nucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00884
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00888
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006177 | GMP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00888
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003922 | GMP synthase (glutamine-hydrolyzing) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00888
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd01742 | GATase1_GMP_Synthase | IPR004739 | GMP synthase, glutamine amidotransferase | 10 | 198 | 1.27603E-97 |
| FunFam | G3DSA:3.30.300.10:FF:000002 | GMP synthase [glutamine-hydrolyzing] | - | - | 411 | 525 | 4.8E-54 |
| SUPERFAMILY | SSF52402 | Adenine nucleotide alpha hydrolases-like | - | - | 197 | 420 | 5.95E-58 |
| Pfam | PF02540 | NAD synthase | IPR022310 | NAD/GMP synthase | 215 | 289 | 2.2E-7 |
| NCBIfam | TIGR00888 | JCVI: glutamine-hydrolyzing GMP synthase, N-terminal domain | IPR004739 | GMP synthase, glutamine amidotransferase | 10 | 205 | 9.3E-78 |
| CDD | cd01997 | GMP_synthase_C | IPR001674 | GMP synthase, C-terminal | 230 | 524 | 0.0 |
| PRINTS | PR00097 | Anthranilate synthase component II signature | - | - | 177 | 190 | 6.6E-6 |
| FunFam | G3DSA:3.40.50.880:FF:000001 | GMP synthase [glutamine-hydrolyzing] | - | - | 7 | 206 | 2.3E-86 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | - | - | 54 | 63 | 4.1E-7 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 209 | 410 | 3.2E-78 |
| NCBIfam | TIGR00884 | JCVI: glutamine-hydrolyzing GMP synthase, C-terminal domain | IPR001674 | GMP synthase, C-terminal | 212 | 525 | 0.0 |
| PRINTS | PR00097 | Anthranilate synthase component II signature | - | - | 81 | 92 | 6.6E-6 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | - | - | 81 | 92 | 4.1E-7 |
| Gene3D | G3DSA:3.40.50.880 | - | IPR029062 | Class I glutamine amidotransferase-like | 9 | 206 | 4.7E-52 |
| Hamap | MF_00344 | GMP synthase [glutamine-hydrolyzing] [guaA]. | IPR022955 | GMP synthase | 7 | 525 | 55.73851 |
| PRINTS | PR00097 | Anthranilate synthase component II signature | - | - | 54 | 63 | 6.6E-6 |
| Gene3D | G3DSA:3.30.300.10 | - | - | - | 411 | 525 | 1.2E-48 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | - | - | 177 | 190 | 4.1E-7 |
| Pfam | PF00958 | GMP synthase C terminal domain | IPR001674 | GMP synthase, C-terminal | 433 | 524 | 1.6E-40 |
| Pfam | PF00117 | Glutamine amidotransferase class-I | IPR017926 | Glutamine amidotransferase | 12 | 198 | 1.6E-35 |
| PANTHER | PTHR11922 | GMP SYNTHASE-RELATED | - | - | 10 | 525 | 0.0 |
| SUPERFAMILY | SSF52317 | Class I glutamine amidotransferase-like | IPR029062 | Class I glutamine amidotransferase-like | 5 | 204 | 1.06E-54 |
| SUPERFAMILY | SSF54810 | GMP synthetase C-terminal dimerisation domain | - | - | 405 | 525 | 2.09E-49 |
| FunFam | G3DSA:3.40.50.620:FF:000001 | GMP synthase [glutamine-hydrolyzing] | - | - | 209 | 410 | 4.0E-101 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.