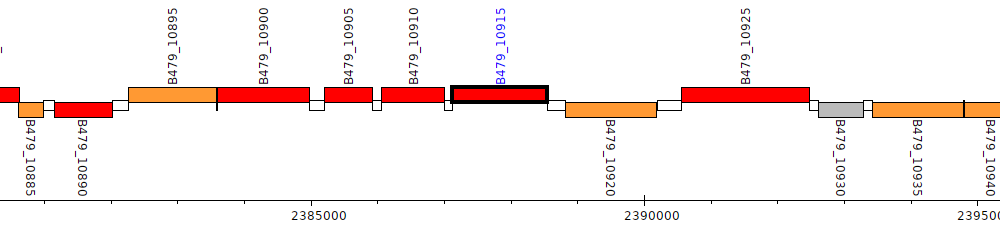
Pseudomonas putida HB3267, B479_10915
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016831 | carboxy-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00800
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016830 | carbon-carbon lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00282
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00800
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00282
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019752 | carboxylic acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00282
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.90.1150.10 | Aspartate Aminotransferase, domain 1 | IPR015422 | Pyridoxal phosphate-dependent transferase, small domain | 353 | 470 | 3.7E-24 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 94 | 112 | 8.3E-46 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 385 | 404 | 8.3E-46 |
| Gene3D | G3DSA:3.40.640.10 | - | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 87 | 352 | 2.0E-88 |
| Pfam | PF00282 | Pyridoxal-dependent decarboxylase conserved domain | IPR002129 | Pyridoxal phosphate-dependent decarboxylase | 35 | 402 | 4.1E-96 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 342 | 357 | 8.3E-46 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 113 | 132 | 8.3E-46 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 6 | 25 | 8.3E-46 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 73 | 92 | 8.3E-46 |
| SUPERFAMILY | SSF53383 | PLP-dependent transferases | IPR015424 | Pyridoxal phosphate-dependent transferase | 1 | 469 | 0.0 |
| PANTHER | PTHR11999 | GROUP II PYRIDOXAL-5-PHOSPHATE DECARBOXYLASE | - | - | 1 | 468 | 0.0 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 29 | 46 | 8.3E-46 |
| Gene3D | G3DSA:1.20.1340.10 | - | - | - | 1 | 81 | 2.9E-33 |
| PRINTS | PR00800 | Aromatic-L-amino-acid decarboxylase signature | IPR010977 | Aromatic-L-amino-acid decarboxylase | 47 | 66 | 8.3E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.