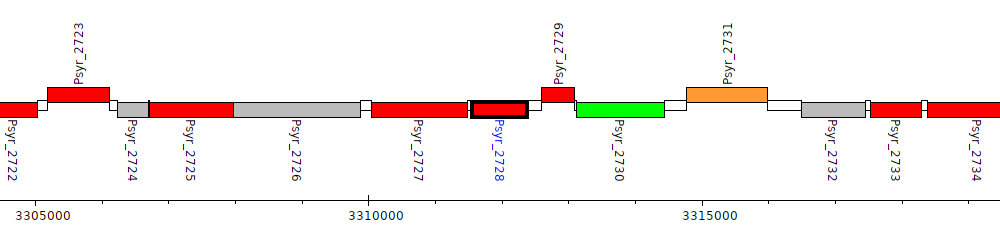
Pseudomonas syringae pv. syringae B728a, Psyr_2728
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psb01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb00360 | Phenylalanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06558 | crotonase-like | - | - | 11 | 208 | 4.6016E-72 |
| FunFam | G3DSA:3.90.226.10:FF:000036 | p-hydroxycinnamoyl CoA hydratase/lyase | - | - | 1 | 211 | 0.0 |
| Gene3D | G3DSA:3.90.226.10 | - | - | - | 1 | 211 | 7.6E-67 |
| PANTHER | PTHR42964 | ENOYL-COA HYDRATASE | - | - | 9 | 266 | 5.2E-65 |
| Gene3D | G3DSA:6.10.250.2850 | - | - | - | 212 | 250 | 6.1E-25 |
| Pfam | PF00378 | Enoyl-CoA hydratase/isomerase | IPR001753 | Enoyl-CoA hydratase/isomerase | 14 | 263 | 5.5E-55 |
| SUPERFAMILY | SSF52096 | ClpP/crotonase | IPR029045 | ClpP/crotonase-like domain superfamily | 8 | 264 | 7.09E-67 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.