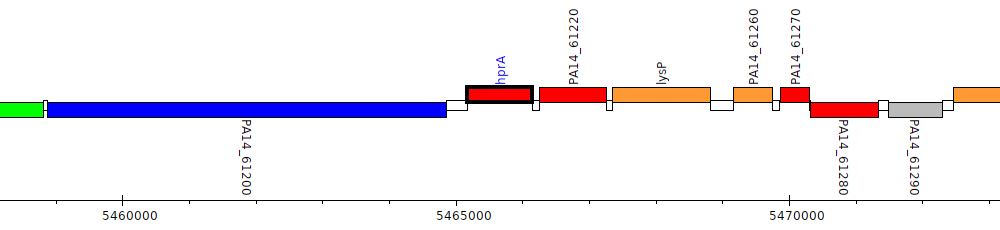
Pseudomonas aeruginosa UCBPP-PA14, PA14_61210 (hprA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00680 | Methane metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43761 | D-ISOMER SPECIFIC 2-HYDROXYACID DEHYDROGENASE FAMILY PROTEIN (AFU_ORTHOLOGUE AFUA_1G13630) | - | - | 1 | 321 | 3.4E-113 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 20 | 317 | 1.6E-111 |
| Pfam | PF02826 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain | IPR006140 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD-binding domain | 112 | 290 | 1.7E-51 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 106 | 291 | 3.2E-52 |
| Pfam | PF00389 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | IPR006139 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | 19 | 320 | 2.1E-25 |
| CDD | cd12162 | 2-Hacid_dh_4 | - | - | 5 | 311 | 0.0 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 102 | 289 | 1.6E-111 |
| SUPERFAMILY | SSF52283 | Formate/glycerate dehydrogenase catalytic domain-like | - | - | 17 | 141 | 3.97E-29 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.