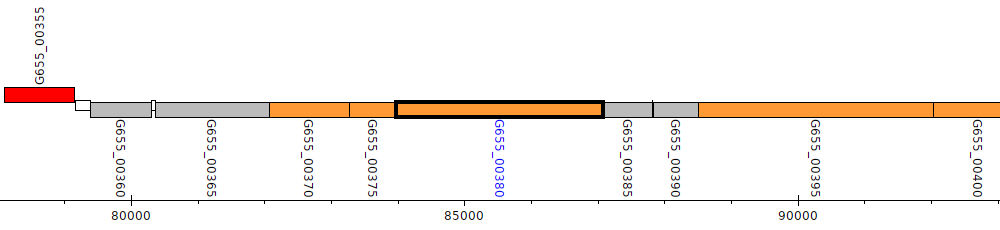
Pseudomonas aeruginosa B136-33, G655_00380
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004674 | protein serine/threonine kinase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0074
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
9852028 | |
| Biological Process | GO:0050714 | positive regulation of protein secretion |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0074
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
19400797 | |
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006468 | protein phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004672 | protein kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00069
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psg02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psg03070 | Bacterial secretion system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd14014 | STKc_PknB_like | - | - | 7 | 261 | 7.69345E-108 |
| PANTHER | PTHR43289 | MITOGEN-ACTIVATED PROTEIN KINASE KINASE KINASE 20-RELATED | - | - | 5 | 407 | 7.8E-66 |
| Gene3D | G3DSA:1.10.510.10 | Transferase(Phosphotransferase) domain 1 | - | - | 88 | 274 | 1.8E-48 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 276 | 350 | - |
| FunFam | G3DSA:1.10.510.10:FF:000021 | Serine/threonine protein kinase | - | - | 89 | 269 | 1.6E-55 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 281 | 299 | - |
| SMART | SM00220 | serkin_6 | IPR000719 | Protein kinase domain | 8 | 263 | 5.1E-66 |
| CDD | cd00198 | vWFA | - | - | 616 | 797 | 2.92108E-12 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 381 | 417 | - |
| SUPERFAMILY | SSF56112 | Protein kinase-like (PK-like) | IPR011009 | Protein kinase-like domain superfamily | 6 | 339 | 8.5E-77 |
| SUPERFAMILY | SSF53300 | vWA-like | IPR036465 | von Willebrand factor A-like domain superfamily | 615 | 793 | 2.23E-12 |
| Gene3D | G3DSA:3.40.50.410 | von Willebrand factor, type A domain | IPR036465 | von Willebrand factor A-like domain superfamily | 614 | 769 | 5.9E-12 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 316 | 347 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 585 | 606 | - |
| SMART | SM00327 | VWA_4 | IPR002035 | von Willebrand factor, type A | 613 | 804 | 2.5E-9 |
| Pfam | PF00092 | von Willebrand factor type A domain | IPR002035 | von Willebrand factor, type A | 616 | 780 | 1.5E-5 |
| Pfam | PF00069 | Protein kinase domain | IPR000719 | Protein kinase domain | 9 | 250 | 3.9E-50 |
| Gene3D | G3DSA:3.30.200.20 | Phosphorylase Kinase; domain 1 | - | - | 2 | 87 | 3.6E-29 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.