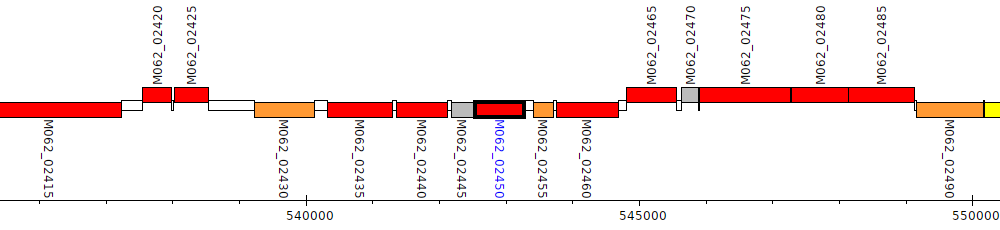
Pseudomonas aeruginosa RP73, M062_02450
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00156 | Phosphoribosyl transferase domain | IPR000836 | Phosphoribosyltransferase domain | 139 | 238 | 7.7E-8 |
| PANTHER | PTHR47505 | DNA UTILIZATION PROTEIN YHGH | - | - | 7 | 238 | 1.4E-66 |
| Gene3D | G3DSA:3.40.50.2020 | - | IPR029057 | Phosphoribosyltransferase-like | 89 | 241 | 5.3E-16 |
| Pfam | PF18912 | Double zinc ribbon domain | IPR044005 | Double zinc ribbon domain | 9 | 65 | 8.6E-9 |
| SUPERFAMILY | SSF53271 | PRTase-like | IPR029057 | Phosphoribosyltransferase-like | 47 | 239 | 1.31E-32 |
| CDD | cd06223 | PRTases_typeI | IPR000836 | Phosphoribosyltransferase domain | 103 | 241 | 1.11237E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.