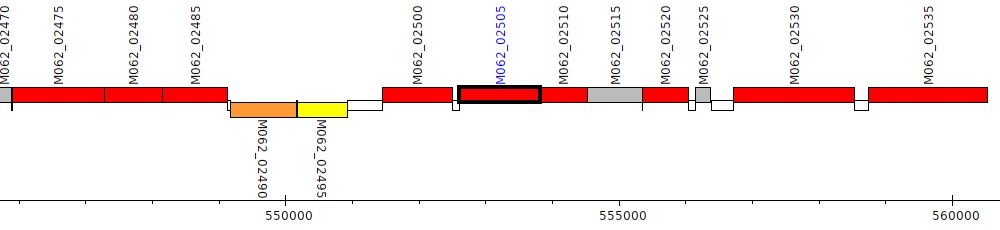
Pseudomonas aeruginosa RP73, M062_02505
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009102 | biotin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00858
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008710 | 8-amino-7-oxononanoate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00858
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | prp00780 | Biotin metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 8-amino-7-oxononanoate biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 8-amino-7-oxononanoate biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | prp01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 8-amino-7-oxononanoate biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06454 | KBL_like | - | - | 37 | 382 | 0.0 |
| Pfam | PF00155 | Aminotransferase class I and II | IPR004839 | Aminotransferase, class I/classII | 39 | 378 | 3.8E-52 |
| NCBIfam | TIGR00858 | JCVI: 8-amino-7-oxononanoate synthase | IPR004723 | 8-amino-7-oxononanoate synthase, Archaea/Proteobacteria type | 17 | 378 | 0.0 |
| Gene3D | G3DSA:3.40.640.10 | - | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 55 | 284 | 0.0 |
| Hamap | MF_01693 | 8-amino-7-oxononanoate synthase [bioF]. | IPR022834 | 8-amino-7-oxononanoate synthase, Proteobacteria | 1 | 382 | 51.290154 |
| SUPERFAMILY | SSF53383 | PLP-dependent transferases | IPR015424 | Pyridoxal phosphate-dependent transferase | 6 | 385 | 1.16E-109 |
| Gene3D | G3DSA:3.90.1150.10 | Aspartate Aminotransferase, domain 1 | IPR015422 | Pyridoxal phosphate-dependent transferase, small domain | 39 | 376 | 0.0 |
| PANTHER | PTHR13693 | CLASS II AMINOTRANSFERASE/8-AMINO-7-OXONONANOATE SYNTHASE | - | - | 22 | 378 | 1.8E-95 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.