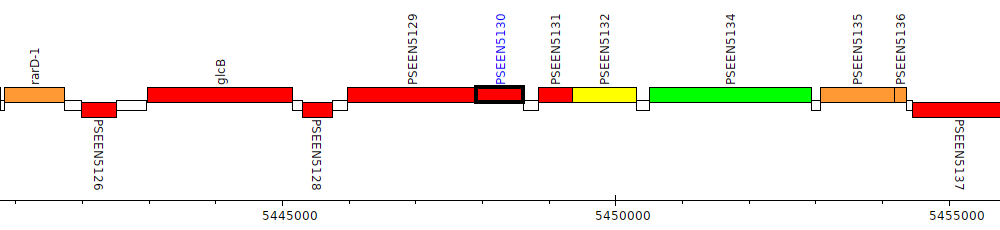
Pseudomonas entomophila L48, PSEEN5130
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.420.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pen00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen03030 | DNA replication | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen03440 | Homologous recombination | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen03430 | Mismatch repair | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pen01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06127 | DEDDh | - | - | 42 | 202 | 1.85004E-36 |
| Pfam | PF00929 | Exonuclease | IPR013520 | Exonuclease, RNase T/DNA polymerase III | 42 | 200 | 5.6E-12 |
| Gene3D | G3DSA:3.30.420.10 | - | IPR036397 | Ribonuclease H superfamily | 32 | 214 | 1.8E-29 |
| PANTHER | PTHR30231 | DNA POLYMERASE III SUBUNIT EPSILON | - | - | 36 | 223 | 1.6E-15 |
| SMART | SM00479 | exoiiiendus | IPR013520 | Exonuclease, RNase T/DNA polymerase III | 40 | 210 | 4.9E-15 |
| SUPERFAMILY | SSF53098 | Ribonuclease H-like | IPR012337 | Ribonuclease H-like superfamily | 35 | 210 | 3.56E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.