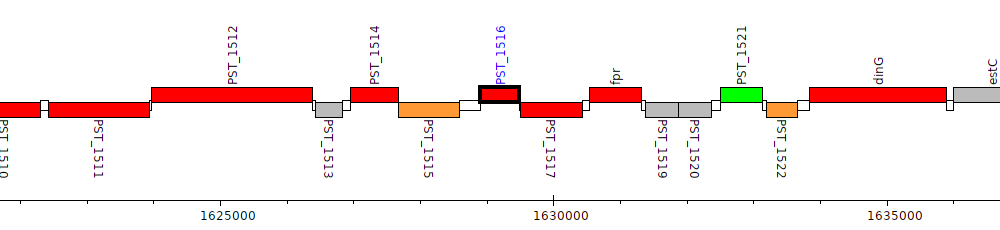
Pseudomonas stutzeri A1501, PST_1516
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46894
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:SM00421
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:PF00072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR45566 | HTH-TYPE TRANSCRIPTIONAL REGULATOR YHJB-RELATED | - | - | 7 | 187 | 4.5E-52 |
| CDD | cd06170 | LuxR_C_like | IPR000792 | Transcription regulator LuxR, C-terminal | 129 | 182 | 2.88742E-17 |
| PRINTS | PR00038 | LuxR bacterial regulatory protein HTH signature | IPR000792 | Transcription regulator LuxR, C-terminal | 129 | 143 | 5.0E-11 |
| CDD | cd17535 | REC_NarL-like | - | - | 11 | 100 | 1.45561E-25 |
| SMART | SM00448 | REC_2 | IPR001789 | Signal transduction response regulator, receiver domain | 1 | 95 | 0.0079 |
| SUPERFAMILY | SSF46894 | C-terminal effector domain of the bipartite response regulators | IPR016032 | Signal transduction response regulator, C-terminal effector | 117 | 189 | 1.02E-20 |
| SMART | SM00421 | luxrmega5 | IPR000792 | Transcription regulator LuxR, C-terminal | 126 | 183 | 1.6E-23 |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 10 | 95 | 2.9E-14 |
| Gene3D | G3DSA:3.40.50.2300 | - | - | - | 4 | 188 | 1.8E-42 |
| PRINTS | PR00038 | LuxR bacterial regulatory protein HTH signature | IPR000792 | Transcription regulator LuxR, C-terminal | 159 | 171 | 5.0E-11 |
| SUPERFAMILY | SSF52172 | CheY-like | IPR011006 | CheY-like superfamily | 6 | 156 | 3.56E-20 |
| PRINTS | PR00038 | LuxR bacterial regulatory protein HTH signature | IPR000792 | Transcription regulator LuxR, C-terminal | 143 | 159 | 5.0E-11 |
| Pfam | PF00196 | Bacterial regulatory proteins, luxR family | IPR000792 | Transcription regulator LuxR, C-terminal | 128 | 182 | 6.6E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.