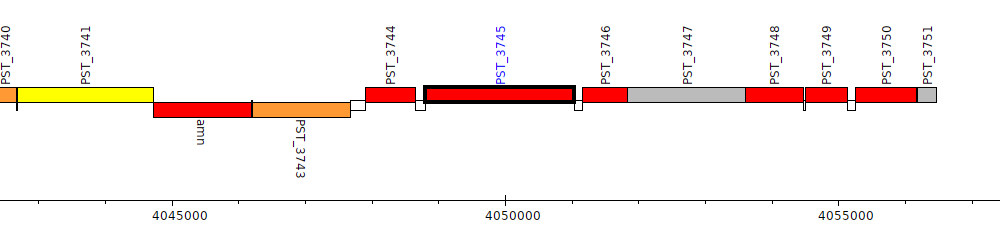
Pseudomonas stutzeri A1501, PST_3745
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00487 | ultradead3 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 222 | 435 | 6.1E-5 |
| NCBIfam | TIGR01596 | JCVI: CRISPR-associated endonuclease Cas3'' | IPR006483 | CRISPR-associated Cas3-type HD domain | 24 | 182 | 2.2E-30 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 21 | - |
| CDD | cd17930 | DEXHc_cas3 | - | - | 242 | 408 | 2.59828E-58 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 436 | 603 | 4.4E-11 |
| SUPERFAMILY | SSF109604 | HD-domain/PDEase-like | - | - | 17 | 181 | 1.23E-13 |
| Pfam | PF01966 | HD domain | IPR006674 | HD domain | 26 | 106 | 2.1E-6 |
| Gene3D | G3DSA:1.10.3210.30 | - | IPR038257 | CRISPR-associated Cas3-type HD domain superfamily | 15 | 133 | 3.9E-10 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 233 | 415 | 1.5E-15 |
| Pfam | PF00270 | DEAD/DEAH box helicase | IPR011545 | DEAD/DEAH box helicase domain | 248 | 406 | 3.6E-8 |
| CDD | cd09641 | Cas3''_I | - | - | 17 | 187 | 3.77408E-33 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 246 | 556 | 1.29E-31 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.