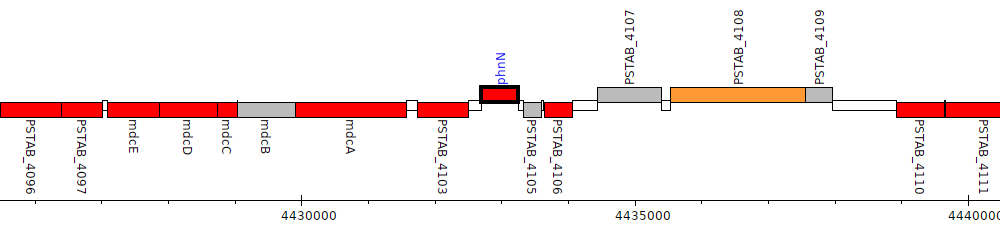
Pseudomonas stutzeri ATCC 17588 = LMG 11199, PSTAB_4104 (phnN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006015 | 5-phosphoribose 1-diphosphate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0033863 | ribose 1,5-bisphosphate phosphokinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00836
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psz00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 183 | 8.1E-26 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 2 | 180 | 1.57E-38 |
| SMART | SM00072 | gk_7 | IPR008145 | Guanylate kinase/L-type calcium channel beta subunit | 2 | 181 | 8.2E-7 |
| Hamap | MF_00836 | Ribose 1,5-bisphosphate phosphokinase PhnN [phnN]. | IPR012699 | Ribose 1,5-bisphosphate phosphokinase PhnN | 2 | 178 | 35.915798 |
| PANTHER | PTHR23117 | GUANYLATE KINASE-RELATED | - | - | 3 | 178 | 1.1E-11 |
| NCBIfam | TIGR02322 | JCVI: phosphonate metabolism protein/1,5-bisphosphokinase (PRPP-forming) PhnN | IPR012699 | Ribose 1,5-bisphosphate phosphokinase PhnN | 3 | 179 | 8.7E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.