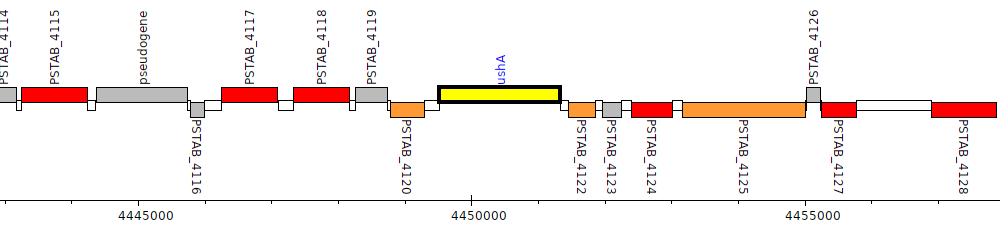
Pseudomonas stutzeri ATCC 17588 = LMG 11199, PSTAB_4121 (ushA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009166 | nucleotide catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR01607
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR01607
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psz01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz00760 | Nicotinate and nicotinamide metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR11575 | 5'-NUCLEOTIDASE-RELATED | IPR006179 | 5'-Nucleotidase/apyrase | 64 | 142 | 2.8E-98 |
| FunFam | G3DSA:3.90.780.10:FF:000004 | UDP-sugar hydrolase, putative | - | - | 417 | 604 | 3.9E-51 |
| Pfam | PF02872 | 5'-nucleotidase, C-terminal domain | IPR008334 | 5'-Nucleotidase, C-terminal | 428 | 567 | 1.5E-42 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 541 | 560 | 2.1E-18 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 69 | 87 | 2.1E-18 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 23 | 59 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 32 | 59 | - |
| SUPERFAMILY | SSF56300 | Metallo-dependent phosphatases | IPR029052 | Metallo-dependent phosphatase-like | 64 | 414 | 9.68E-56 |
| Pfam | PF00149 | Calcineurin-like phosphoesterase | IPR004843 | Calcineurin-like phosphoesterase domain, ApaH type | 72 | 334 | 8.2E-9 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 281 | 298 | 2.1E-18 |
| SUPERFAMILY | SSF55816 | 5'-nucleotidase (syn. UDP-sugar hydrolase), C-terminal domain | IPR036907 | 5'-Nucleotidase, C-terminal domain superfamily | 415 | 604 | 4.45E-50 |
| PRINTS | PR01607 | Apyrase family signature | IPR006179 | 5'-Nucleotidase/apyrase | 469 | 492 | 2.1E-18 |
| Gene3D | G3DSA:3.90.780.10 | - | IPR036907 | 5'-Nucleotidase, C-terminal domain superfamily | 423 | 604 | 1.8E-50 |
| Gene3D | G3DSA:3.60.21.10 | - | IPR029052 | Metallo-dependent phosphatase-like | 66 | 414 | 1.1E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.