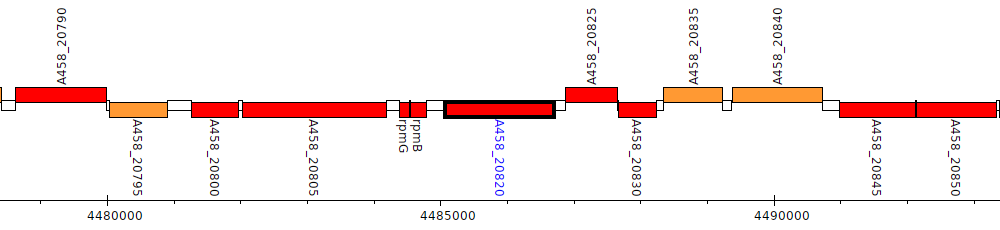
Pseudomonas stutzeri CCUG 29243, A458_20820
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psc02024 | Quorum sensing | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psc00061 | Fatty acid biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psc01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psc00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psc01212 | Fatty acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00501 | AMP-binding enzyme | IPR000873 | AMP-dependent synthetase/ligase domain | 26 | 443 | 4.8E-103 |
| PANTHER | PTHR43767 | LONG-CHAIN-FATTY-ACID--COA LIGASE | - | - | 19 | 532 | 1.1E-125 |
| FunFam | G3DSA:3.40.50.12780:FF:000003 | Long-chain-fatty-acid--CoA ligase FadD | - | - | 9 | 438 | 9.0E-115 |
| Pfam | PF13193 | AMP-binding enzyme C-terminal domain | IPR025110 | AMP-binding enzyme, C-terminal domain | 452 | 526 | 6.7E-20 |
| Gene3D | G3DSA:3.30.300.30 | - | IPR045851 | AMP-binding enzyme, C-terminal domain superfamily | 439 | 539 | 4.5E-30 |
| SUPERFAMILY | SSF56801 | Acetyl-CoA synthetase-like | - | - | 14 | 538 | 0.0 |
| CDD | cd05936 | FC-FACS_FadD_like | - | - | 23 | 533 | 0.0 |
| Gene3D | G3DSA:3.40.50.12780 | - | IPR042099 | ANL, N-terminal domain | 11 | 438 | 1.2E-114 |
| FunFam | G3DSA:3.30.300.30:FF:000006 | Long-chain-fatty-acid--CoA ligase FadD | - | - | 438 | 539 | 1.1E-42 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.