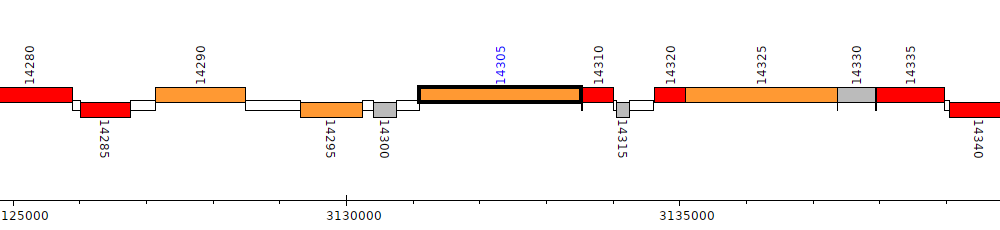
Pseudomonas stutzeri DSM 10701, PSJM300_14305
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00003
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005215 | transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 612 | 623 | 2.4E-29 |
| Gene3D | G3DSA:3.40.1110.10 | - | IPR023299 | P-type ATPase, cytoplasmic domain N | 499 | 618 | 1.8E-31 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 455 | 744 | 1.1E-58 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 1 | 55 | 4.9E-12 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 409 | 423 | 2.8E-10 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 710 | 722 | 2.4E-29 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 258 | 765 | 3.6E-92 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 62 | 122 | 6.81053E-19 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 59 | 125 | 1.7E-20 |
| FunFam | G3DSA:2.70.150.10:FF:000020 | Copper-exporting P-type ATPase A | - | - | 274 | 390 | 2.8E-42 |
| Gene3D | G3DSA:2.70.150.10 | - | - | - | 274 | 390 | 5.9E-39 |
| Pfam | PF00122 | E1-E2 ATPase | - | - | 287 | 467 | 2.4E-57 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 687 | 706 | 2.4E-29 |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | - | - | 470 | 738 | 0.0 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 634 | 644 | 2.4E-29 |
| SUPERFAMILY | SSF81653 | Calcium ATPase, transduction domain A | IPR008250 | P-type ATPase, A domain superfamily | 290 | 386 | 7.59E-30 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 58 | 130 | 2.5E-20 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 486 | 701 | 2.2E-44 |
| NCBIfam | TIGR00003 | JCVI: copper ion binding protein | IPR006122 | Heavy metal-associated domain, copper ion-binding | 1 | 56 | 8.2E-10 |
| NCBIfam | TIGR01525 | JCVI: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 219 | 796 | 0.0 |
| SFLD | SFLDF00027 | p-type atpase | IPR044492 | P-type ATPase, haloacid dehalogenase domain | 470 | 738 | 0.0 |
| FunFam | G3DSA:3.30.70.100:FF:000005 | Copper-exporting P-type ATPase A | - | - | 57 | 124 | 4.9E-22 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 336 | 350 | 2.4E-29 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 1 | 57 | 4.0E-17 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 760 | 774 | 2.8E-10 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 1 | 56 | 2.92987E-14 |
| SUPERFAMILY | SSF81665 | Calcium ATPase, transmembrane domain M | IPR023298 | P-type ATPase, transmembrane domain superfamily | 237 | 767 | 2.09E-12 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 486 | 794 | 8.25E-62 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 63 | 120 | 1.5E-16 |
| CDD | cd02094 | P-type_ATPase_Cu-like | - | - | 147 | 798 | 0.0 |
| FunFam | G3DSA:3.40.1110.10:FF:000099 | Copper-translocating P-type ATPase | - | - | 499 | 620 | 4.0E-52 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 664 | 681 | 2.8E-10 |
| PANTHER | PTHR43520 | ATP7, ISOFORM B | - | - | 1 | 800 | 0.0 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 488 | 502 | 2.4E-29 |
| NCBIfam | TIGR01511 | JCVI: copper-translocating P-type ATPase | - | - | 201 | 797 | 0.0 |
| Coils | Coil | Coil | - | - | 128 | 148 | - |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 256 | 275 | 2.8E-10 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 204 | 223 | 2.8E-10 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 1 | 56 | 3.53E-14 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.