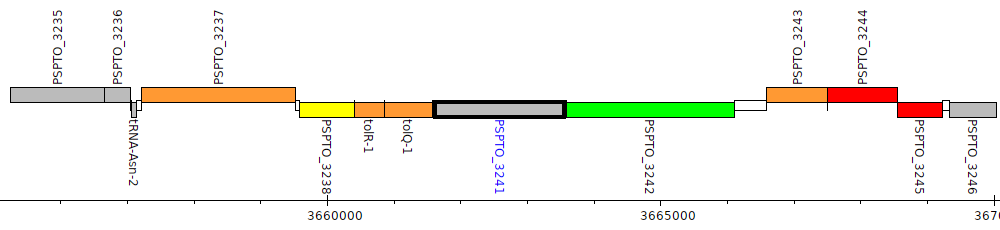
Pseudomonas syringae pv. tomato DC3000, PSPTO_3241
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016158 | 3-phytase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02333
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pst00562 | Inositol phosphate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF50956 | Thermostable phytase (3-phytase) | - | - | 14 | 310 | 5.36E-38 |
| SUPERFAMILY | SSF50956 | Thermostable phytase (3-phytase) | - | - | 319 | 640 | 6.41E-82 |
| Pfam | PF02333 | Phytase | IPR003431 | Beta propeller phytase | 323 | 637 | 3.3E-78 |
| Gene3D | G3DSA:2.120.10.30 | - | IPR011042 | Six-bladed beta-propeller, TolB-like | 52 | 273 | 3.8E-24 |
| Gene3D | G3DSA:2.120.10.30 | - | IPR011042 | Six-bladed beta-propeller, TolB-like | 316 | 640 | 8.8E-118 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.