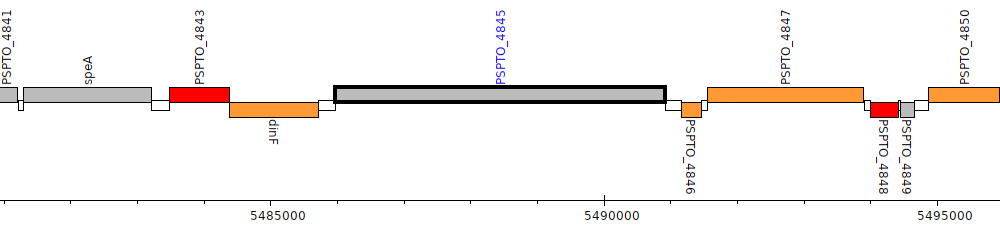
Pseudomonas syringae pv. tomato DC3000, PSPTO_4845
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004866 | endopeptidase inhibitor activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM01360
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005615 | extracellular space |
Inferred from Sequence Model
Term mapped from: InterPro:PF07678
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | Coil | - | - | 1550 | 1584 | - |
| SMART | SM01360 | A2M_2 | IPR001599 | Alpha-2-macroglobulin | 963 | 1052 | 5.5E-24 |
| Pfam | PF07703 | Alpha-2-macroglobulin bait region domain | IPR011625 | Alpha-2-macroglobulin, bait region domain | 755 | 901 | 2.1E-10 |
| Pfam | PF01835 | MG2 domain | IPR002890 | Macroglobulin domain | 382 | 474 | 1.4E-20 |
| PANTHER | PTHR40094 | ALPHA-2-MACROGLOBULIN HOMOLOG | - | - | 127 | 1640 | 0.0 |
| SUPERFAMILY | SSF48239 | Terpenoid cyclases/Protein prenyltransferases | IPR008930 | Terpenoid cyclases/protein prenyltransferase alpha-alpha toroid | 1170 | 1426 | 1.47E-38 |
| Pfam | PF00207 | Alpha-2-macroglobulin family | IPR001599 | Alpha-2-macroglobulin | 972 | 1051 | 3.1E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 22 | 42 | - |
| SMART | SM01419 | Thiol_ester_cl_2 | IPR047565 | Alpha-macroglobulin-like, thiol-ester bond-forming region | 1171 | 1200 | 4.1E-5 |
| Pfam | PF17973 | Bacterial Alpha-2-macroglobulin MG10 domain | IPR041246 | Bacterial alpha-2-macroglobulin MG10 domain | 1500 | 1632 | 1.6E-25 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 26 | 40 | - |
| Pfam | PF17972 | Bacterial Alpha-2-macroglobulin MG5 domain | IPR041203 | Bacterial Alpha-2-macroglobulin, MG5 domain | 480 | 605 | 4.8E-41 |
| SMART | SM01359 | A2M_N_2_2 | IPR011625 | Alpha-2-macroglobulin, bait region domain | 754 | 902 | 2.9E-38 |
| Pfam | PF17970 | Bacterial Alpha-2-macroglobulin MG1 domain | IPR040639 | Alpha-2-macroglobulin, MG1 domain | 54 | 157 | 5.6E-51 |
| Gene3D | G3DSA:2.60.40.1930 | - | - | - | 381 | 478 | 6.7E-15 |
| Pfam | PF11974 | Bacterial alpha-2-macroglobulin MG3 domain | IPR021868 | Alpha-2-macroglobulin MG3 domain | 283 | 376 | 5.4E-23 |
| Pfam | PF07678 | A-macroglobulin TED domain | IPR011626 | Alpha-macroglobulin-like, TED domain | 1170 | 1292 | 3.1E-10 |
| PIRSF | PIRSF038980 | A2M_bac | IPR026284 | Alpha-2-macroglobulin, bacteria | 2 | 1649 | 0.0 |
| CDD | cd02891 | A2M_like | - | - | 1170 | 1430 | 6.54718E-53 |
| Pfam | PF17962 | Bacterial macroglobulin domain 6 | IPR041462 | Bacterial Alpha-2-macroglobulin, MG6 domain | 633 | 738 | 3.2E-30 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 615 | 639 | - |
| Gene3D | G3DSA:1.50.10.20 | - | - | - | 1167 | 1423 | 2.5E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.