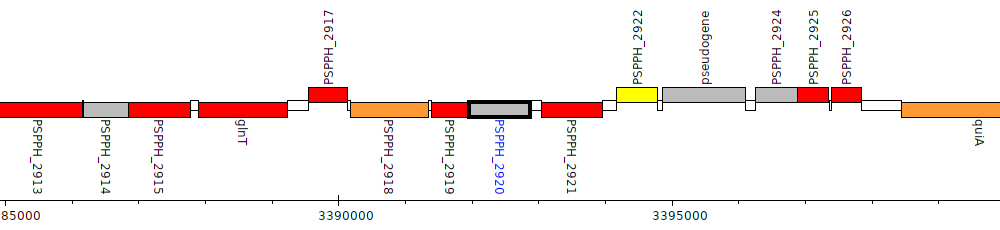
Pseudomonas savastanoi pv. phaseolicola 1448A, PSPPH_2920
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003933 | GTP cyclohydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR36445
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003934 | GTP cyclohydrolase I activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01527_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0035998 | 7,8-dihydroneopterin 3'-triphosphate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01527_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psp00790 | Folate biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.10.270.10 | Urate Oxidase; | - | - | 3 | 298 | 8.4E-63 |
| PANTHER | PTHR36445 | GTP CYCLOHYDROLASE MPTA | IPR003801 | GTP cyclohydrolase FolE2/MptA | 3 | 297 | 2.7E-67 |
| Hamap | MF_01527_B | GTP cyclohydrolase FolE2 [folE2]. | IPR022838 | GTP cyclohydrolase FolE2 | 5 | 299 | 33.546665 |
| Pfam | PF02649 | Type I GTP cyclohydrolase folE2 | IPR003801 | GTP cyclohydrolase FolE2/MptA | 5 | 294 | 4.0E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.