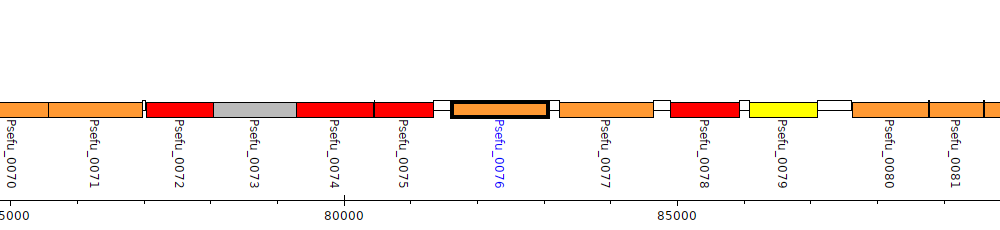
Pseudomonas fulva 12-X, Psefu_0076
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008808 | cardiolipin synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04265
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0032049 | cardiolipin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04265
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04265
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pfv01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfv00564 | Glycerophospholipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR04265 | JCVI: cardiolipin synthase | IPR022924 | Cardiolipin synthase | 18 | 479 | 4.3E-112 |
| Gene3D | G3DSA:3.30.870.10 | Endonuclease Chain A | - | - | 114 | 289 | 1.5E-48 |
| Gene3D | G3DSA:3.30.870.10 | Endonuclease Chain A | - | - | 301 | 453 | 4.5E-27 |
| Hamap | MF_00190 | Cardiolipin synthase A [clsA]. | IPR030840 | Cardiolipin synthase A | 2 | 479 | 48.349762 |
| Pfam | PF13091 | PLD-like domain | IPR025202 | Phospholipase D-like domain | 320 | 441 | 6.6E-27 |
| PANTHER | PTHR21248 | CARDIOLIPIN SYNTHASE | - | - | 31 | 479 | 1.3E-101 |
| FunFam | G3DSA:3.30.870.10:FF:000021 | Cardiolipin synthase | - | - | 302 | 451 | 5.9E-66 |
| SUPERFAMILY | SSF56024 | Phospholipase D/nuclease | - | - | 270 | 454 | 2.92E-48 |
| SMART | SM00155 | pld_4 | IPR001736 | Phospholipase D/Transphosphatidylase | 392 | 419 | 1.3E-5 |
| Pfam | PF13091 | PLD-like domain | IPR025202 | Phospholipase D-like domain | 135 | 278 | 1.9E-9 |
| SMART | SM00155 | pld_4 | IPR001736 | Phospholipase D/Transphosphatidylase | 218 | 245 | 3.9E-5 |
| CDD | cd09155 | PLDc_PaCLS_like_1 | - | - | 126 | 281 | 2.32207E-92 |
| FunFam | G3DSA:3.30.870.10:FF:000014 | Cardiolipin synthase | - | - | 113 | 297 | 9.0E-73 |
| SUPERFAMILY | SSF56024 | Phospholipase D/nuclease | - | - | 84 | 308 | 7.22E-56 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.