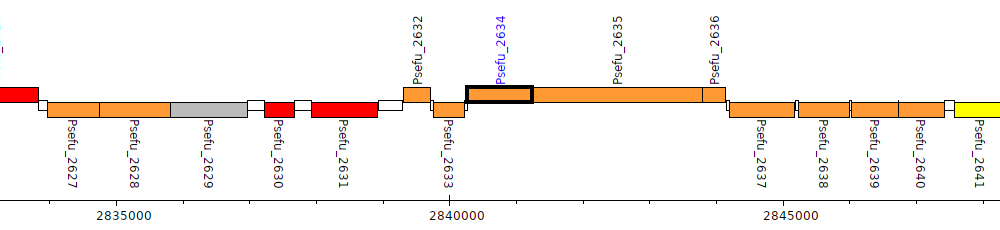
Pseudomonas fulva 12-X, Psefu_2634
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02866
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46626
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46626
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR22888
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02866
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00116
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | arsenite oxidation I (respiratory) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | aerobic respiration I (cytochrome c) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | aerobic respiration II (cytochrome c) (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | Fe(II) oxidation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pfv00190 | Oxidative phosphorylation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfv01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF81464 | Cytochrome c oxidase subunit II-like, transmembrane region | IPR036257 | Cytochrome C oxidase subunit II, transmembrane domain superfamily | 16 | 84 | 6.87E-5 |
| Pfam | PF00116 | Cytochrome C oxidase subunit II, periplasmic domain | IPR002429 | Cytochrome c oxidase subunit II-like C-terminal | 136 | 211 | 8.2E-20 |
| SUPERFAMILY | SSF49503 | Cupredoxins | IPR008972 | Cupredoxin | 110 | 265 | 1.14E-50 |
| CDD | cd04213 | CuRO_CcO_Caa3_II | IPR034236 | Caa3-type Cytochrome c oxidase subunit II, cupredoxin domain | 113 | 212 | 7.81705E-66 |
| SUPERFAMILY | SSF46626 | Cytochrome c | IPR036909 | Cytochrome c-like domain superfamily | 239 | 320 | 1.31E-11 |
| Gene3D | G3DSA:2.60.40.420 | - | IPR008972 | Cupredoxin | 111 | 322 | 4.0E-73 |
| NCBIfam | TIGR02866 | JCVI: cytochrome c oxidase subunit II | IPR014222 | Cytochrome c oxidase, subunit II | 23 | 221 | 5.3E-56 |
| Gene3D | G3DSA:1.10.287.90 | - | IPR036257 | Cytochrome C oxidase subunit II, transmembrane domain superfamily | 16 | 106 | 6.5E-7 |
| PANTHER | PTHR22888 | CYTOCHROME C OXIDASE, SUBUNIT II | IPR045187 | Cytochrome c/quinol oxidase subunit II | 5 | 229 | 7.0E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.