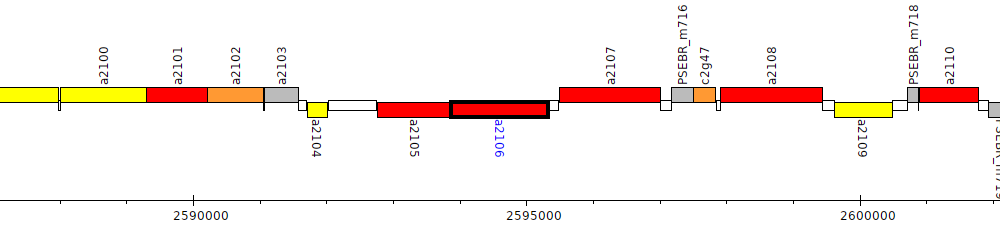
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a2106
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005506 | iron ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016966 | nitric oxide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01312
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005243
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF005243 | ROO | IPR016440 | Rubredoxin-oxygen oxidoreductase | 1 | 399 | 0.0 |
| PRINTS | PR00163 | Rubredoxin signature | IPR024935 | Rubredoxin domain | 426 | 442 | 3.6E-5 |
| Gene3D | G3DSA:2.20.28.10 | - | - | - | 424 | 480 | 3.2E-18 |
| Pfam | PF00753 | Metallo-beta-lactamase superfamily | IPR001279 | Metallo-beta-lactamase | 34 | 170 | 1.8E-9 |
| CDD | cd07709 | flavodiiron_proteins_MBL-fold | - | - | 3 | 246 | 1.45761E-110 |
| Gene3D | G3DSA:3.60.15.10 | - | IPR036866 | Ribonuclease Z/Hydroxyacylglutathione hydrolase-like | 1 | 247 | 7.9E-109 |
| Hamap | MF_01312 | Anaerobic nitric oxide reductase flavorubredoxin [norV]. | IPR023957 | Anaerobic nitric oxide reductase flavorubredoxin | 1 | 476 | 84.289642 |
| Gene3D | G3DSA:3.40.50.360 | - | IPR029039 | Flavoprotein-like superfamily | 249 | 399 | 1.9E-40 |
| Pfam | PF00258 | Flavodoxin | IPR008254 | Flavodoxin/nitric oxide synthase | 256 | 386 | 1.1E-10 |
| PANTHER | PTHR43717 | ANAEROBIC NITRIC OXIDE REDUCTASE FLAVORUBREDOXIN | - | - | 2 | 395 | 0.0 |
| SUPERFAMILY | SSF57802 | Rubredoxin-like | - | - | 426 | 476 | 3.81E-15 |
| SUPERFAMILY | SSF52218 | Flavoproteins | IPR029039 | Flavoprotein-like superfamily | 251 | 393 | 7.86E-35 |
| CDD | cd00730 | rubredoxin | IPR024935 | Rubredoxin domain | 426 | 475 | 4.23732E-17 |
| SMART | SM00849 | Lactamase_B_5a | IPR001279 | Metallo-beta-lactamase | 34 | 210 | 1.4E-28 |
| SUPERFAMILY | SSF56281 | Metallo-hydrolase/oxidoreductase | IPR036866 | Ribonuclease Z/Hydroxyacylglutathione hydrolase-like | 8 | 285 | 4.06E-47 |
| PRINTS | PR00163 | Rubredoxin signature | IPR024935 | Rubredoxin domain | 456 | 472 | 3.6E-5 |
| Pfam | PF00301 | Rubredoxin | IPR024935 | Rubredoxin domain | 426 | 472 | 1.3E-16 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.