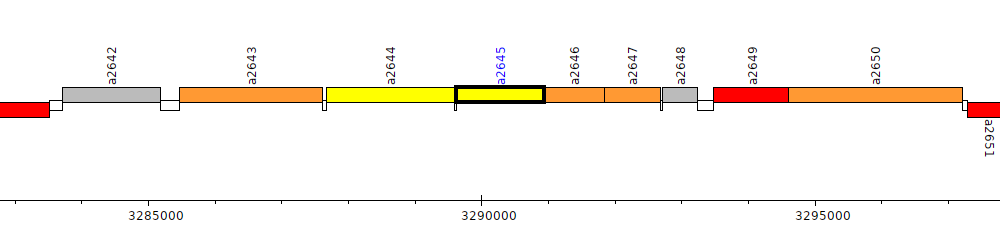
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a2645
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00722 | complete | IPR006633 | Carbohydrate-binding/sugar hydrolysis domain | 41 | 190 | 1.6E-24 |
| Gene3D | G3DSA:2.160.20.10 | - | IPR012334 | Pectin lyase fold | 204 | 358 | 2.0E-12 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 140 | 169 | 1400.0 |
| Pfam | PF05048 | Periplasmic copper-binding protein (NosD) | IPR007742 | Periplasmic copper-binding protein NosD, beta helix domain | 146 | 359 | 3.9E-54 |
| SUPERFAMILY | SSF51126 | Pectin lyase-like | IPR011050 | Pectin lyase fold/virulence factor | 40 | 411 | 1.01E-55 |
| NCBIfam | TIGR03804 | JCVI: parallel beta-helix repeat (two copies) | IPR022441 | Parallel beta-helix repeat-2 | 164 | 203 | 2.3E-7 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 214 | 235 | 5500.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 192 | 213 | 1400.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 236 | 258 | 760.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 295 | 316 | 100.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 118 | 139 | 12.0 |
| NCBIfam | TIGR04247 | JCVI: nitrous oxide reductase family maturation protein NosD | IPR026464 | Nitrous oxide reductase family maturation protein NosD | 41 | 423 | 0.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 170 | 191 | 14.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 318 | 356 | 9700.0 |
| SMART | SM00710 | pbh1 | IPR006626 | Parallel beta-helix repeat | 88 | 116 | 780.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 417 | 439 | - |
| SMART | SM00722 | complete | IPR006633 | Carbohydrate-binding/sugar hydrolysis domain | 196 | 370 | 1.1E-30 |
| Gene3D | G3DSA:2.160.20.10 | - | IPR012334 | Pectin lyase fold | 74 | 203 | 1.0E-10 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.