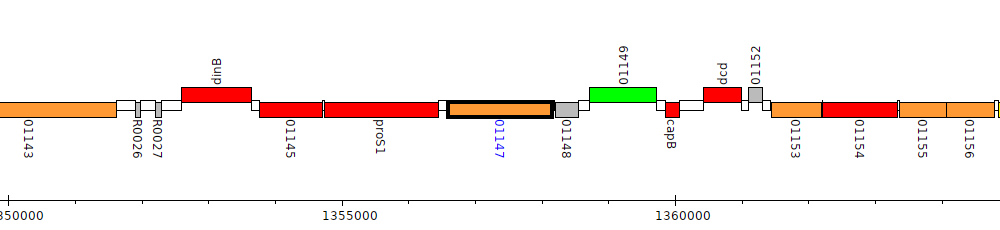
Pseudomonas sp. UW4, PputUW4_01147
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR12778
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuu01501 | beta-Lactam resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07690 | Major Facilitator Superfamily | IPR011701 | Major facilitator superfamily | 21 | 316 | 2.4E-9 |
| SUPERFAMILY | SSF103473 | MFS general substrate transporter | IPR036259 | MFS transporter superfamily | 2 | 497 | 3.92E-31 |
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 10 | 220 | 9.8E-10 |
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 311 | 497 | 1.2E-9 |
| FunFam | G3DSA:1.20.1250.20:FF:000072 | Muropeptide transporter AmpG | - | - | 15 | 219 | 3.2E-75 |
| PANTHER | PTHR12778 | SOLUTE CARRIER FAMILY 33 ACETYL-COA TRANSPORTER -RELATED | IPR004752 | AmpG-like permease/Acetyl-coenzyme A transporter 1 | 4 | 493 | 5.5E-75 |
| NCBIfam | TIGR00901 | JCVI: AmpG family muropeptide MFS transporter | - | - | 29 | 455 | 1.6E-129 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.