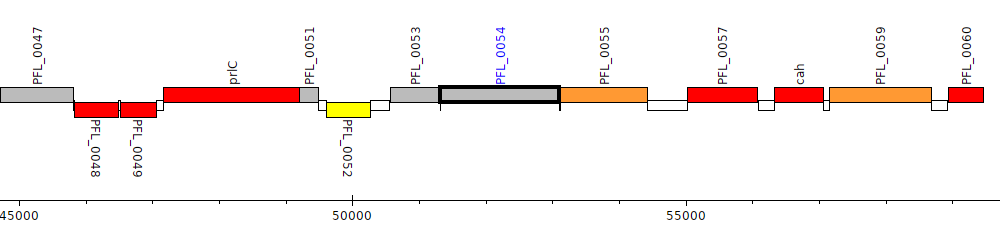
Pseudomonas protegens Pf-5, PFL_0054
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00732
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016614 | oxidoreductase activity, acting on CH-OH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:PF00732
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pfl00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfl01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfl01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 475 | 586 | 1.8E-7 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 2 | 372 | 1.5E-27 |
| Pfam | PF05199 | GMC oxidoreductase | IPR007867 | Glucose-methanol-choline oxidoreductase, C-terminal | 451 | 573 | 7.6E-21 |
| SUPERFAMILY | SSF54373 | FAD-linked reductases, C-terminal domain | - | - | 433 | 496 | 6.67E-10 |
| Pfam | PF00732 | GMC oxidoreductase | IPR000172 | Glucose-methanol-choline oxidoreductase, N-terminal | 203 | 348 | 1.1E-7 |
| PANTHER | PTHR46056 | LONG-CHAIN-ALCOHOL OXIDASE | - | - | 5 | 587 | 1.2E-55 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 6 | 585 | 1.38E-49 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.