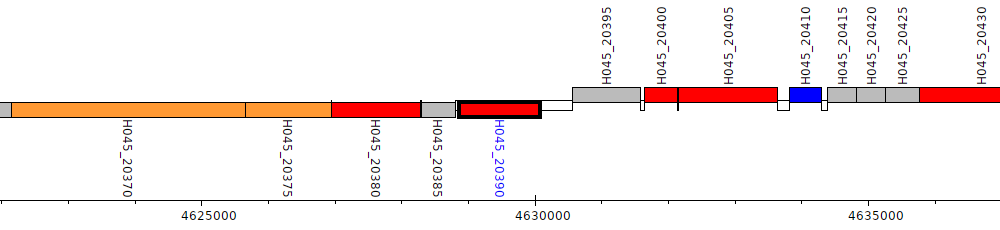
Pseudomonas poae RE_1-1-14, H045_20390
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00498
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppz03070 | Bacterial secretion system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppz02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF49879 | SMAD/FHA domain | IPR008984 | SMAD/FHA domain superfamily | 2 | 88 | 8.03E-21 |
| Gene3D | G3DSA:2.60.200.20 | - | - | - | 1 | 89 | 1.6E-19 |
| SMART | SM00240 | FHA_2 | IPR000253 | Forkhead-associated (FHA) domain | 1 | 51 | 0.0043 |
| NCBIfam | TIGR03354 | JCVI: type VI secretion system-associated FHA domain protein TagH | IPR017735 | Type VI secretion system, FHA domain-containing protein | 2 | 402 | 8.3E-119 |
| Pfam | PF00498 | FHA domain | IPR000253 | Forkhead-associated (FHA) domain | 3 | 70 | 5.9E-15 |
| Pfam | PF20232 | C-terminal domain of Type VI secretion system FHA protein | IPR046883 | Type VI secretion system, FHA domain-containing protein, C-terminal domain | 222 | 400 | 4.6E-64 |
| CDD | cd00060 | FHA | - | - | 3 | 75 | 2.78289E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.