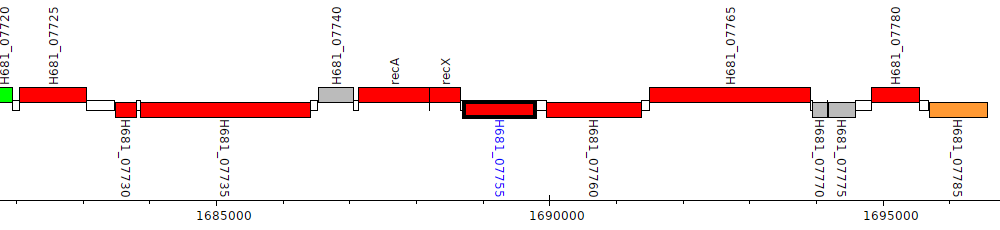
Pseudomonas denitrificans ATCC 13867, H681_07755
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009691 | cytokinin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.40.50.450:FF:000027 | Possible lysine decarboxylase | - | - | 62 | 318 | 1.1E-130 |
| Pfam | PF03641 | Possible lysine decarboxylase | IPR031100 | LOG family | 130 | 263 | 4.5E-30 |
| SUPERFAMILY | SSF102405 | MCP/YpsA-like | - | - | 68 | 263 | 9.62E-49 |
| NCBIfam | TIGR00730 | JCVI: Rossman fold protein, TIGR00730 family | IPR005269 | Cytokinin riboside 5'-monophosphate phosphoribohydrolase LOG | 87 | 265 | 6.4E-28 |
| Gene3D | G3DSA:3.40.50.450 | - | - | - | 63 | 280 | 3.3E-56 |
| PANTHER | PTHR43393 | CYTOKININ RIBOSIDE 5'-MONOPHOSPHATE PHOSPHORIBOHYDROLASE | - | - | 38 | 280 | 1.9E-75 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.