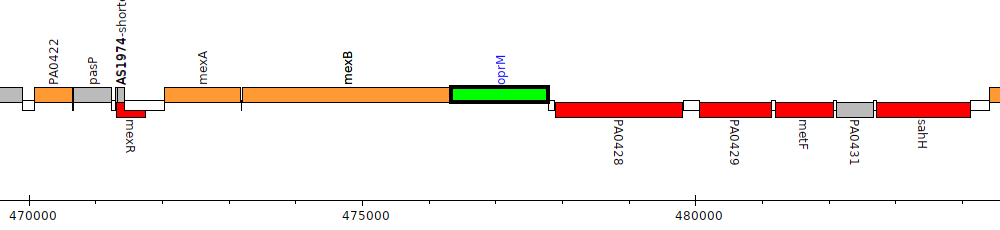
Pseudomonas aeruginosa PAO1, PA0427 (oprM)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0427
|
| Name |
oprM
|
| Replicon | chromosome |
| Genomic location | 476333 - 477790 (+ strand) |
| Transposon Mutants | 1 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
| Comment | Localication to outer membrane vesicle confirmed in three separate studies (PMID:22909304, PMID:21751344 and PMID:26303878). |
Cross-References
| RefSeq | NP_249118.1 |
| GI | 15595624 |
| Affymetrix | PA0427_oprM_at |
| Entrez | 877851 |
| GenBank | AAG03816.1 |
| INSDC | AAA74438.1 |
| NCBI Locus Tag | PA0427 |
| protein_id(GenBank) | gb|AAG03816.1|AE004479_11|gnl|PseudoCAP|PA0427 |
| TIGR | NTL03PA00428 |
| UniParc | UPI0000130D9F |
| UniProtKB Acc | Q51487 |
| UniProtKB ID | OPRM_PSEAE |
| UniRef100 | UniRef100_Q51487 |
| UniRef50 | UniRef50_Q51487 |
| UniRef90 | UniRef90_Q51487 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
Major intrinsic multiple antibiotic resistance efflux outer membrane protein OprM precursor
|
| Synonyms | |
| Evidence for Translation |
Detected using LC-MS/MS (52.60 kDa, PMID:21751344)
Detected by MALDI-TOF/TOF (PMID:19333994).
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -2.79 |
| Kyte-Doolittle Hydrophobicity Value | -0.215 |
| Molecular Weight (kDa) | 52.6 |
| Isoelectric Point (pI) | 5.32 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AMR gene predictions from CARD database
Identified using the Resistance Gene Identifier (RGI)
| Model Name | Definition | Accession | Model Type (?) | SNP | Cutoff | Bit Score | Percent Identity | CARD Version |
| OprM | OprM is an outer membrane factor protein found in Pseudomonas aeruginosa and Burkholderia vietnamiensis. It is part of the MexAB-OprM, MexVW-OprM, MexXY-OprM and the AmrAB-OprM complex. [PMID:16508113, PMID:16511029, PMID:15797729] | ARO:3000379 | protein homolog model | Perfect | 916.8 | 100.0 | 3.2.5 |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7AKZ | TRANSLOCASE | 10/02/20 | Deciphering the role of the channel constrictions in the opening mechanism of MexAB-OprM efflux pump from Pseudomonas aeruginosa | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 3.2 | X-RAY DIFFRACTION | 100.0 |
| 6TA6 | ANTIMICROBIAL PROTEIN | 10/29/19 | MexAB assembly of the Pseudomonas MexAB-OprM efflux pump reconstituted in nanodiscs | Pseudomonas aeruginosa | 3.2 | ELECTRON MICROSCOPY | 100.0 |
| 1WP1 | MEMBRANE PROTEIN | 08/28/04 | Crystal structure of the drug-discharge outer membrane protein, OprM | Pseudomonas aeruginosa | 2.56 | X-RAY DIFFRACTION | 100.0 |
| 6IOL | MEMBRANE PROTEIN | 10/30/18 | Cryo-EM structure of multidrug efflux pump MexAB-OprM (60 degree state) | Pseudomonas aeruginosa; Pseudomonas aeruginosa PAO1 | 3.76 | ELECTRON MICROSCOPY | 100.0 |
| 4Y1K | MEMBRANE PROTEIN | 02/07/15 | PALMITOYLATED OPRM OUTER MEMBRANE FACTOR | Pseudomonas aeruginosa | 3.8 | X-RAY DIFFRACTION | 100.0 |
| 6IOK | MEMBRANE PROTEIN | 10/30/18 | Cryo-EM structure of multidrug efflux pump MexAB-OprM (0 degree state) | Pseudomonas aeruginosa PAO1 | 3.64 | ELECTRON MICROSCOPY | 100.0 |
| 6ZRE | TRANSLOCASE | 07/13/20 | Deciphering the role of the channel constrictions in the opening mechanism of MexAB-OprM efflux pump from Pseudomonas aeruginosa | Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) | 2.8 | X-RAY DIFFRACTION | 100.0 |
| 6TA5 | ANTIMICROBIAL PROTEIN | 10/29/19 | OprM-MexA complex from the MexAB-OprM Pseudomonas aeruginosa whole assembly reconstituted in nanodiscs | Pseudomonas aeruginosa | 3.2 | ELECTRON MICROSCOPY | 100.0 |
| 3D5K | MEMBRANE PROTEIN | 05/16/08 | Crystal structure of the OprM channel in a non-symmetrical space group | Pseudomonas aeruginosa | 2.4 | X-RAY DIFFRACTION | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 316 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000418 (534 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0427
Search term: oprM
|
Human Homologs
References
|
Evidence for the location of OprM in the Pseudomonas aeruginosa outer membrane.
Gotoh N, Itoh N, Yamada H, Nishino T
FEMS Microbiol Lett 1994 Oct 1;122(3):309-12
PubMed ID: 7988873
|
|
Multiple antibiotic resistance in Pseudomonas aeruginosa: evidence for involvement of an efflux operon.
Poole K, Krebes K, McNally C, Neshat S
J Bacteriol 1993 Nov;175(22):7363-72
PubMed ID: 8226684
|
|
Role of mexA-mexB-oprM in antibiotic efflux in Pseudomonas aeruginosa.
Li XZ, Nikaido H, Poole K
Antimicrob Agents Chemother 1995 Sep;39(9):1948-53
PubMed ID: 8540696
|
|
The outer membrane protein OprM of Pseudomonas aeruginosa is encoded by oprK of the mexA-mexB-oprK multidrug resistance operon.
Gotoh N, Tsujimoto H, Poole K, Yamagishi J, Nishino T
Antimicrob Agents Chemother 1995 Nov;39(11):2567-9
PubMed ID: 8585747
|
|
The role of mex-gene products in antibiotic extrusion in Pseudomonas aeruginosa.
Yoneyama H, Ocaktan A, Tsuda M, Nakae T
Biochem Biophys Res Commun 1997 Apr 28;233(3):611-8
PubMed ID: 9168899
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|
|
Proteomic analysis of outer membrane vesicles derived from Pseudomonas aeruginosa.
Choi DS, Kim DK, Choi SJ, Lee J, Choi JP, Rho S, Park SH, Kim YK, Hwang D, Gho YS
Proteomics 2011 Aug;11(16):3424-9
PubMed ID: 21751344
|
|
Identification of proteins associated with the Pseudomonas aeruginosa biofilm extracellular matrix.
Toyofuku M, Roschitzki B, Riedel K, Eberl L
J Proteome Res 2012 Oct 5;11(10):4906-15
PubMed ID: 22909304
|
|
Proteome Profiles of Outer Membrane Vesicles and Extracellular Matrix of Pseudomonas aeruginosa Biofilms.
Couto N, Schooling SR, Dutcher JR, Barber J
J Proteome Res 2015 Oct 2;14(10):4207-22
PubMed ID: 26303878
|