Pseudomonas aeruginosa PAO1, PA1498 (pykF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
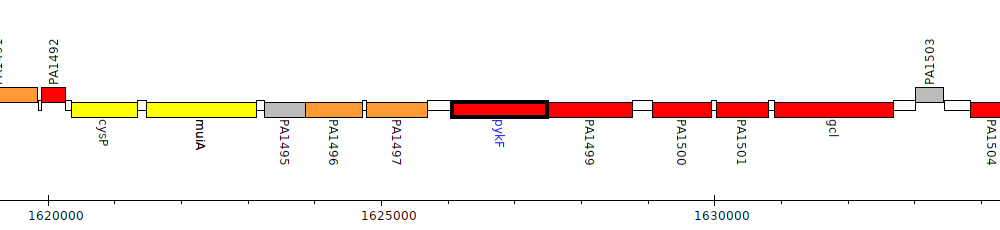
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA1498
|
| Name |
pykF
Synonym: pyk-I |
| Replicon | chromosome |
| Genomic location | 1626062 - 1627495 (- strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_250189.1 |
| GI | 15596695 |
| Affymetrix | PA1498_pykF_at |
| Entrez | 881682 |
| GenBank | AAG04887.1 |
| INSDC | AAG04887.1 |
| NCBI Locus Tag | PA1498 |
| protein_id(GenBank) | gb|AAG04887.1|AE004578_9|gnl|PseudoCAP|PA1498 |
| TIGR | NTL03PA01499 |
| UniParc | UPI00000C53AC |
| UniProtKB Acc | Q9I3L4 |
| UniProtKB ID | Q9I3L4_PSEAE |
| UniRef100 | UniRef100_Q9I3L4 |
| UniRef50 | UniRef50_Q44473 |
| UniRef90 | UniRef90_Q9I3L4 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
pyruvate kinase I
|
| Synonyms |
Phosphoenolpyruvate kinase |
| Evidence for Translation | |
| Charge (pH 7) | -6.41 |
| Kyte-Doolittle Hydrophobicity Value | -0.048 |
| Molecular Weight (kDa) | 51.5 |
| Isoelectric Point (pI) | 5.56 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7OO1 | TRANSFERASE | 05/26/21 | Structure, function and characterization of a second pyruvate kinase isozyme in Pseudomonas aeruginosa. | Pseudomonas aeruginosa PAO1 | 3.01 | X-RAY DIFFRACTION | 100.0 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 614 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001412 (474 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA1498
Search term: pykF
Search term: pyruvate kinase I
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
pyruvate kinase M1/2 [Source:HGNC Symbol;Acc:HGNC:9021]
E-value:
1.3e-78
Percent Identity:
39.6
|
|
Ensembl
110, assembly
GRCh38.p14
|
pyruvate kinase M1/2 [Source:HGNC Symbol;Acc:HGNC:9021]
E-value:
1.3e-78
Percent Identity:
39.6
|
References
|
Primary structure of three peptides at the catalytic and allosteric sites of the fructose-1,6-bisphosphate-activated pyruvate kinase from Escherichia coli.
Speranza ML, Valentini G, Iadarola P, Stoppini M, Malcovati M, Ferri G
Biol Chem Hoppe Seyler 1989 Mar;370(3):211-6
PubMed ID: 2653362
|
|
Key residues in the allosteric transition of Bacillus stearothermophilus pyruvate kinase identified by site-directed mutagenesis.
Walker D, Chia WN, Muirhead H
J Mol Biol 1992 Nov 5;228(1):265-76
PubMed ID: 1447787
|
|
Crystal structure of Escherichia coli pyruvate kinase type I: molecular basis of the allosteric transition.
Mattevi A, Valentini G, Rizzi M, Speranza ML, Bolognesi M, Coda A
Structure 1995 Jul 15;3(7):729-41
PubMed ID: 8591049
|