Pseudomonas aeruginosa PAO1, PA4117 (bphP)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
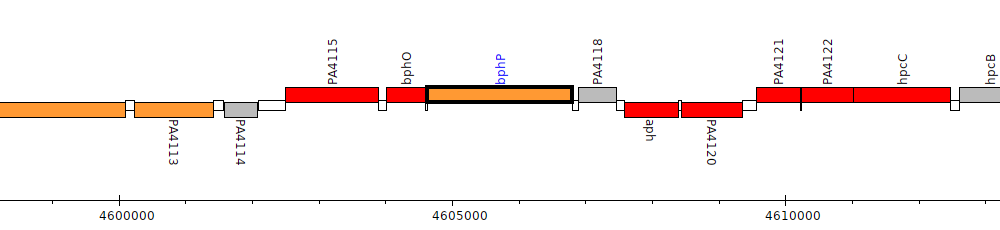
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA4117
|
| Name |
bphP
|
| Replicon | chromosome |
| Genomic location | 4604621 - 4606807 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_252806.1 |
| GI | 15599312 |
| Affymetrix | PA4117_at |
| Entrez | 880365 |
| GenBank | AAG07504.1 |
| INSDC | AAG07504.1 |
| NCBI Locus Tag | PA4117 |
| protein_id(GenBank) | gb|AAG07504.1|AE004828_5|gnl|PseudoCAP|PA4117 |
| TIGR | NTL03PA04117 |
| UniParc | UPI0000126A88 |
| UniProtKB Acc | Q9HWR3 |
| UniProtKB ID | BPHY_PSEAE |
| UniRef100 | UniRef100_Q9HWR3 |
| UniRef50 | UniRef50_Q9HWR3 |
| UniRef90 | UniRef90_Q9HWR3 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
bacterial phytochrome, BphP
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -15.27 |
| Kyte-Doolittle Hydrophobicity Value | -0.200 |
| Molecular Weight (kDa) | 80.9 |
| Isoelectric Point (pI) | 5.70 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 3C2W | SIGNALING PROTEIN | 01/25/08 | Crystal structure of the photosensory core domain of P. aeruginosa bacteriophytochrome PaBphP in the Pfr state | Pseudomonas aeruginosa | 2.9 | X-RAY DIFFRACTION | 99.8 |
| 3NOU | SIGNALING PROTEIN | 06/25/10 | Light-induced intermediate structure L3 of P. aeruginosa bacteriophytochrome | Pseudomonas aeruginosa | 3 | X-RAY DIFFRACTION | 99.8 |
| 3IBR | TRANSFERASE | 07/16/09 | Crystal Structure of P. aeruginosa Bacteriophytochrome Photosensory Core Module Mutant Q188L in the Mixed Pr/Pfr State | Pseudomonas aeruginosa | 2.97 | X-RAY DIFFRACTION | 99.6 |
| 3NOP | SIGNALING PROTEIN | 06/25/10 | Light-induced intermediate structure L1 of Pseudomonas aeruginosa bacteriophytochrome | Pseudomonas aeruginosa | 2.8 | X-RAY DIFFRACTION | 99.8 |
| 3NHQ | SIGNALING PROTEIN | 06/14/10 | The dark Pfr structure of the photosensory core module of P. aeruginosa Bacteriophytochrome | Pseudomonas aeruginosa | 2.55 | X-RAY DIFFRACTION | 99.8 |
| 3G6O | SIGNALING PROTEIN | 02/07/09 | Crystal structure of P. aeruginosa bacteriophytochrome PaBphP photosensory core domain mutant Q188L | Pseudomonas aeruginosa | 2.85 | X-RAY DIFFRACTION | 99.6 |
| 3NOT | SIGNALING PROTEIN | 06/25/10 | Light-induced intermediate structure L2 of P. aeruginosa bacteriophytochrome | Pseudomonas Aeruginosa | 2.7 | X-RAY DIFFRACTION | 99.8 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 615 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001581 (502 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA4117
Search term: bphP
Search term: bacterial phytochrome, BphP
|
Human Homologs
References
|
Sequence and characterization of a novel chromosomal aminoglycoside phosphotransferase gene, aph (3')-IIb, in Pseudomonas aeruginosa.
Hächler H, Santanam P, Kayser FH
Antimicrob Agents Chemother 1996 May;40(5):1254-6
PubMed ID: 8723476
|