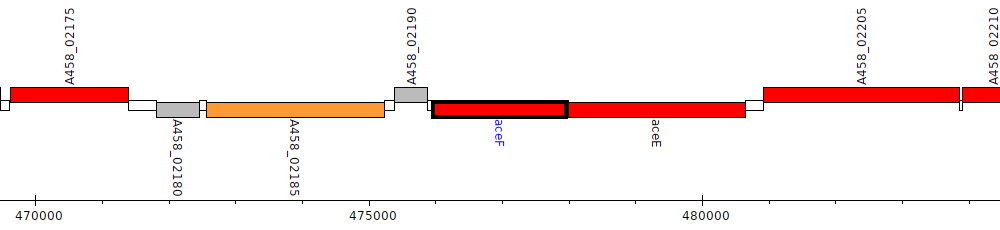
Pseudomonas stutzeri CCUG 29243, A458_02195 (aceF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas stutzeri CCUG 29243
GCF_000267545.1|latest |
| Locus Tag |
A458_02195
|
| Name |
aceF
|
| Replicon | chromosome |
| Genomic location | 475961 - 477967 (- strand) |
Cross-References
| RefSeq | YP_006456118.1 |
| GI | 392419514 |
| EC Number | 2.3.1.12 |
| Entrez | 13145219 |
| INSDC | AFM31695.1 |
| NCBI Locus Tag | A458_02195 |
| UniParc | UPI0002660C8E |
| UniProtKB Acc | I4CNN8 |
| UniProtKB ID | I4CNN8_PSEST |
| UniRef100 | UniRef100_I4CNN8 |
| UniRef50 | UniRef50_P10802 |
| UniRef90 | UniRef90_F2N4Q1 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
pyruvate dehydrogenase dihydrolipoyltransacetylase
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -23.07 |
| Kyte-Doolittle Hydrophobicity Value | -0.112 |
| Molecular Weight (kDa) | 69068.9 |
| Isoelectric Point (pI) | 4.69 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 1DPD | DIHYDROLIPOAMIDE ACETYLTRANSFERASE | 02/03/95 | CRYSTALLOGRAPHIC AND ENZYMATIC INVESTIGATIONS ON THE ROLE OF SER558, HIS610 AND ASN614 IN THE CATALYTIC MECHANISM OF AZOTOBACTER VINELANDII DIHYDROLIPOAMIDE ACETYLTRANSFERASE (E2P) | Azotobacter vinelandii | 2.7 | X-RAY DIFFRACTION | 90.1 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 524 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG004623 (531 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: A458_02195
Search term: aceF
Search term: pyruvate dehydrogenase dihydrolipoyltransacetylase
|
Human Homologs
|
Ensembl
110, assembly
GRCh38.p14
|
dihydrolipoamide S-acetyltransferase [Source:HGNC Symbol;Acc:HGNC:2896]
E-value:
1.1e-33
Percent Identity:
33.2
|
|
Ensembl
110, assembly
GRCh38.p14
|
dihydrolipoamide S-acetyltransferase [Source:HGNC Symbol;Acc:HGNC:2896]
E-value:
1.1e-33
Percent Identity:
33.2
|
|
Ensembl
110, assembly
GRCh38.p14
|
dihydrolipoamide S-acetyltransferase [Source:HGNC Symbol;Acc:HGNC:2896]
E-value:
1.1e-33
Percent Identity:
33.2
|
|
Ensembl
110, assembly
GRCh38.p14
|
dihydrolipoamide S-acetyltransferase [Source:HGNC Symbol;Acc:HGNC:2896]
E-value:
1.1e-33
Percent Identity:
33.2
|
|
Ensembl
110, assembly
GRCh38.p14
|
dihydrolipoamide S-acetyltransferase [Source:HGNC Symbol;Acc:HGNC:2896]
E-value:
1.1e-33
Percent Identity:
33.2
|
References
No references are associated with this feature.