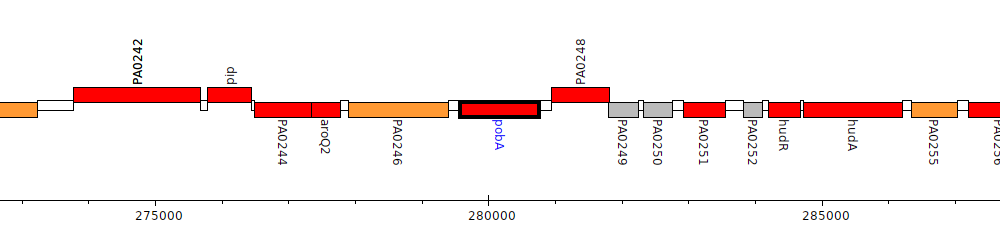
Pseudomonas aeruginosa PAO1, PA0247 (pobA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0018659 | 4-hydroxybenzoate 3-monooxygenase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2465205 | Reviewed by curator |
| Biological Process | GO:0043639 | benzoate catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2465205 | Reviewed by curator |
| Molecular Function | GO:0018659 | 4-hydroxybenzoate 3-monooxygenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02360
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0043639 | benzoate catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02360
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02360
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0071949 | FAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00362 | Benzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Aromatic compound catabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | 4-HYDROXYMANDELATE-DEGRADATION-PWY | 4-hydroxymandelate degradation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01220 | Degradation of aromatic compounds | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01494 | FAD binding domain | IPR002938 | FAD-binding domain | 2 | 343 | 3.0E-100 |
| PRINTS | PR00420 | Aromatic-ring hydroxylase (flavoprotein monooxygenase) signature | - | - | 310 | 328 | 1.5E-32 |
| Gene3D | G3DSA:3.30.9.10 | - | - | - | 73 | 388 | 0.0 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 389 | 9.43E-59 |
| PRINTS | PR00420 | Aromatic-ring hydroxylase (flavoprotein monooxygenase) signature | - | - | 293 | 309 | 1.5E-32 |
| SUPERFAMILY | SSF54373 | FAD-linked reductases, C-terminal domain | - | - | 175 | 275 | 2.47E-46 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 2 | 351 | 0.0 |
| PANTHER | PTHR43476 | 3-(3-HYDROXY-PHENYL)PROPIONATE/3-HYDROXYCINNAMIC ACID HYDROXYLASE | - | - | 3 | 328 | 6.8E-31 |
| PRINTS | PR00420 | Aromatic-ring hydroxylase (flavoprotein monooxygenase) signature | - | - | 151 | 166 | 1.5E-32 |
| PRINTS | PR00420 | Aromatic-ring hydroxylase (flavoprotein monooxygenase) signature | - | - | 278 | 293 | 1.5E-32 |
| PRINTS | PR00420 | Aromatic-ring hydroxylase (flavoprotein monooxygenase) signature | - | - | 4 | 26 | 1.5E-32 |
| NCBIfam | TIGR02360 | JCVI: 4-hydroxybenzoate 3-monooxygenase | IPR012733 | 4-hydroxybenzoate 3-monooxygenase | 1 | 390 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.