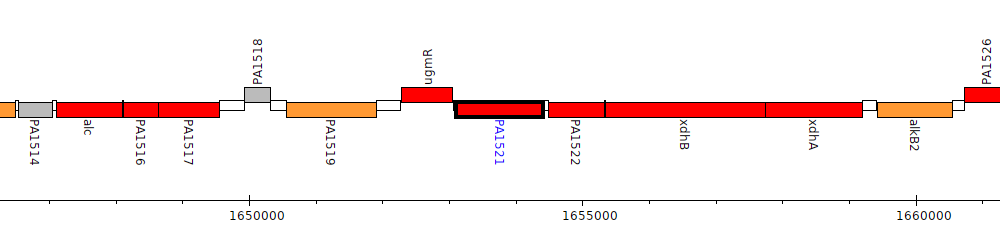
Pseudomonas aeruginosa PAO1, PA1521
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008892 | guanine deaminase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd01303
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd01303
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006147 | guanine catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd01303
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.30.40.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01979
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Purine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.30.40.10 | Urease, subunit C, domain 1 | IPR011059 | Metal-dependent hydrolase, composite domain superfamily | 9 | 424 | 0.0 |
| Pfam | PF01979 | Amidohydrolase family | IPR006680 | Amidohydrolase-related | 69 | 430 | 4.3E-69 |
| Gene3D | G3DSA:3.20.20.140 | - | - | - | 72 | 390 | 0.0 |
| NCBIfam | TIGR02967 | JCVI: guanine deaminase | IPR014311 | Guanine deaminase | 28 | 427 | 0.0 |
| FunFam | G3DSA:3.20.20.140:FF:000022 | Guanine deaminase | - | - | 72 | 373 | 0.0 |
| SUPERFAMILY | SSF51338 | Composite domain of metallo-dependent hydrolases | IPR011059 | Metal-dependent hydrolase, composite domain superfamily | 29 | 88 | 2.07E-10 |
| SUPERFAMILY | SSF51556 | Metallo-dependent hydrolases | IPR032466 | Metal-dependent hydrolase | 71 | 374 | 4.0E-104 |
| PANTHER | PTHR11271 | GUANINE DEAMINASE | - | - | 3 | 433 | 7.8E-119 |
| CDD | cd01303 | GDEase | IPR014311 | Guanine deaminase | 8 | 427 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.