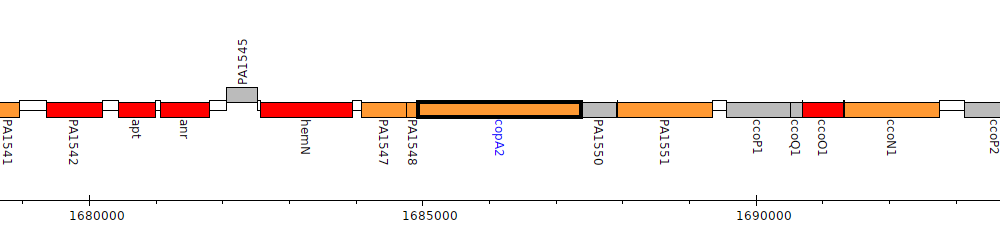
Pseudomonas aeruginosa PAO1, PA1549 (copA2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55008
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005215 | transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43520 | ATP7, ISOFORM B | - | - | 91 | 798 | 0.0 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 108 | 129 | 4.7E-5 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 90 | 159 | 1.9E-12 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 284 | 772 | 9.9E-67 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 506 | 794 | 1.84E-49 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 97 | 156 | 3.5E-8 |
| SUPERFAMILY | SSF81653 | Calcium ATPase, transduction domain A | IPR008250 | P-type ATPase, A domain superfamily | 323 | 409 | 2.09E-22 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 717 | 729 | 5.0E-21 |
| NCBIfam | TIGR01525 | JCVI: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 248 | 796 | 0.0 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 694 | 713 | 5.0E-21 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 360 | 374 | 5.0E-21 |
| FunFam | G3DSA:2.70.150.10:FF:000002 | Copper-transporting ATPase 1, putative | - | - | 297 | 414 | 3.5E-31 |
| Pfam | PF12156 | Putative metal-binding domain of cation transport ATPase | IPR021993 | Putative metal-binding domain of cation transport ATPase | 7 | 89 | 6.9E-28 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 641 | 651 | 5.0E-21 |
| Gene3D | G3DSA:3.40.1110.10 | - | IPR023299 | P-type ATPase, cytoplasmic domain N | 519 | 627 | 2.6E-64 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 508 | 522 | 5.0E-21 |
| Gene3D | G3DSA:2.70.150.10 | - | - | - | 297 | 414 | 7.3E-29 |
| CDD | cd02079 | P-type_ATPase_HM | - | - | 184 | 795 | 0.0 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 504 | 708 | 9.6E-33 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 97 | 106 | 4.7E-5 |
| Pfam | PF00122 | E1-E2 ATPase | - | - | 310 | 487 | 1.7E-48 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 97 | 156 | 8.38441E-9 |
| SUPERFAMILY | SSF81665 | Calcium ATPase, transmembrane domain M | IPR023298 | P-type ATPase, transmembrane domain superfamily | 262 | 501 | 5.36E-8 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 376 | 392 | 4.7E-5 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 501 | 744 | 2.6E-64 |
| NCBIfam | TIGR01511 | JCVI: copper-translocating P-type ATPase | - | - | 231 | 796 | 0.0 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 87 | 164 | 6.68E-17 |
| NCBIfam | TIGR01512 | JCVI: cadmium family heavy metal-translocating P-type ATPase | - | - | 276 | 797 | 2.0E-112 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.