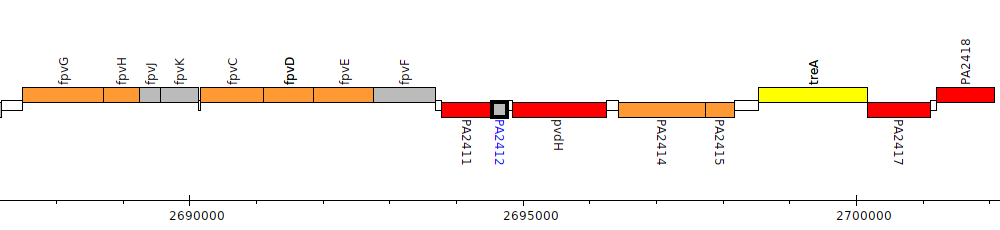
Pseudomonas aeruginosa PAO1, PA2412
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016790 | thiolester hydrolase activity | Inferred from Sequence or Structural Similarity | ECO:0000247 sequence alignment evidence used in manual assertion |
12207696 | Reviewed by curator |
| Biological Process | GO:0002049 | pyoverdine biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
12686626 | Reviewed by curator |
| Biological Process | GO:0002049 | pyoverdine biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
12207696 | Reviewed by curator |
| Biological Process | GO:0002049 | pyoverdine biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
17502378 | Reviewed by curator |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00261 | Monobactam biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00923 | MbtH_2 | IPR005153 | MbtH-like domain | 3 | 53 | 6.0E-27 |
| SUPERFAMILY | SSF160582 | MbtH-like | IPR038020 | MbtH-like domain superfamily | 1 | 69 | 2.48E-30 |
| FunFam | G3DSA:3.90.820.10:FF:000003 | Antibiotic synthesis protein MbtH | - | - | 1 | 72 | 1.7E-54 |
| PANTHER | PTHR38444 | ENTEROBACTIN BIOSYNTHESIS PROTEIN YBDZ | IPR037407 | MbtH-like protein | 1 | 69 | 6.7E-30 |
| Gene3D | G3DSA:3.90.820.10 | - | - | - | 1 | 72 | 4.7E-36 |
| Pfam | PF03621 | MbtH-like protein | IPR005153 | MbtH-like domain | 1 | 53 | 5.5E-25 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.