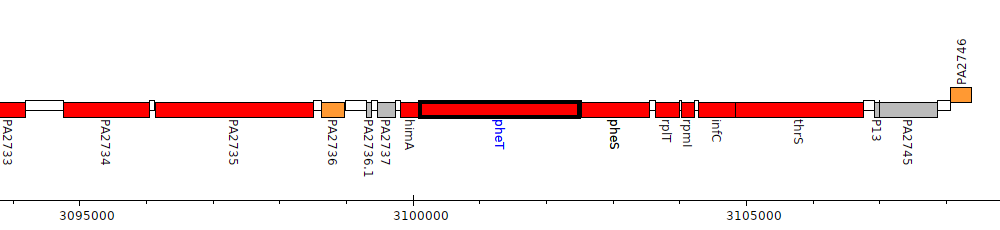
Pseudomonas aeruginosa PAO1, PA2739 (pheT)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006259 | DNA metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0004817 | cysteine-tRNA ligase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00874
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006432 | phenylalanyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR10947
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000049 | tRNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd02796
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00874
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004826 | phenylalanine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR10947
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF03483
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Phenylalanine, tyrosine and tryptophan biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | TRNA-CHARGING-PWY | tRNA charging | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Aminoacyl-tRNA biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.30.930.10:FF:000022 | Phenylalanine--tRNA ligase beta subunit | - | - | 487 | 689 | 3.2E-70 |
| SMART | SM00873 | B3_4_2 | IPR005146 | B3/B4 tRNA-binding domain | 211 | 384 | 2.1E-95 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 39 | 190 | 2.4E-48 |
| FunFam | G3DSA:3.50.40.10:FF:000001 | Phenylalanine--tRNA ligase beta subunit | - | - | 195 | 396 | 1.7E-82 |
| PANTHER | PTHR10947 | PHENYLALANYL-TRNA SYNTHETASE BETA CHAIN AND LEUCINE-RICH REPEAT-CONTAINING PROTEIN 47 | IPR045060 | Phenylalanine-tRNA ligase, class IIc, beta subunit | 2 | 683 | 1.3E-74 |
| SUPERFAMILY | SSF46955 | Putative DNA-binding domain | IPR009061 | Putative DNA-binding domain superfamily | 403 | 475 | 8.33E-18 |
| Hamap | MF_00283 | Phenylalanine--tRNA ligase beta subunit [pheT]. | IPR004532 | Phenylalanine-tRNA ligase, class IIc, beta subunit, bacterial type | 1 | 785 | 46.855408 |
| Gene3D | G3DSA:3.30.930.10 | Bira Bifunctional Protein; Domain 2 | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 485 | 690 | 1.7E-68 |
| CDD | cd00769 | PheRS_beta_core | IPR041616 | Phenylalanyl tRNA synthetase beta chain, core domain | 506 | 683 | 4.51049E-67 |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 39 | 153 | 3.3E-73 |
| SMART | SM00874 | B5_2 | IPR005147 | tRNA synthetase, B5-domain | 402 | 471 | 1.8E-27 |
| Gene3D | G3DSA:3.30.70.380 | - | IPR036690 | Ferrodoxin-fold anticodon-binding domain superfamily | 699 | 792 | 7.7E-28 |
| SUPERFAMILY | SSF56037 | PheT/TilS domain | - | - | 195 | 394 | 1.8E-74 |
| FunFam | G3DSA:2.40.50.140:FF:000045 | Phenylalanine--tRNA ligase beta subunit | - | - | 39 | 153 | 8.6E-47 |
| NCBIfam | TIGR00472 | JCVI: phenylalanine--tRNA ligase subunit beta | IPR004532 | Phenylalanine-tRNA ligase, class IIc, beta subunit, bacterial type | 1 | 791 | 0.0 |
| CDD | cd02796 | tRNA_bind_bactPheRS | IPR033714 | Phenylalanly tRNA synthetase, tRNA-binding-domain | 45 | 146 | 8.81811E-51 |
| Gene3D | G3DSA:3.30.56.10 | - | - | - | 6 | 181 | 3.3E-73 |
| Gene3D | G3DSA:3.30.56.10 | - | - | - | 400 | 475 | 1.6E-22 |
| FunFam | G3DSA:3.30.56.10:FF:000002 | Phenylalanine--tRNA ligase beta subunit | - | - | 401 | 475 | 6.4E-23 |
| Pfam | PF17759 | Phenylalanyl tRNA synthetase beta chain CLM domain | IPR041616 | Phenylalanyl tRNA synthetase beta chain, core domain | 474 | 681 | 3.2E-61 |
| Pfam | PF01588 | Putative tRNA binding domain | IPR002547 | tRNA-binding domain | 45 | 144 | 1.2E-24 |
| FunFam | G3DSA:3.30.70.380:FF:000001 | Phenylalanine--tRNA ligase beta subunit | - | - | 699 | 792 | 5.9E-33 |
| Gene3D | G3DSA:3.50.40.10 | - | IPR020825 | Phenylalanyl-tRNA synthetase-like, B3/B4 | 196 | 394 | 3.6E-80 |
| Pfam | PF03483 | B3/4 domain | IPR005146 | B3/B4 tRNA-binding domain | 211 | 383 | 1.5E-70 |
| SMART | SM00896 | FDX_ACB_2 | IPR005121 | Ferrodoxin-fold anticodon-binding domain | 698 | 791 | 1.9E-46 |
| SUPERFAMILY | SSF55681 | Class II aaRS and biotin synthetases | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 467 | 683 | 8.45E-54 |
| Pfam | PF03484 | tRNA synthetase B5 domain | IPR005147 | tRNA synthetase, B5-domain | 405 | 471 | 1.3E-19 |
| Pfam | PF03147 | Ferredoxin-fold anticodon binding domain | IPR005121 | Ferrodoxin-fold anticodon-binding domain | 698 | 791 | 1.2E-30 |
| SUPERFAMILY | SSF54991 | Anticodon-binding domain of PheRS | IPR036690 | Ferrodoxin-fold anticodon-binding domain superfamily | 692 | 791 | 2.09E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.