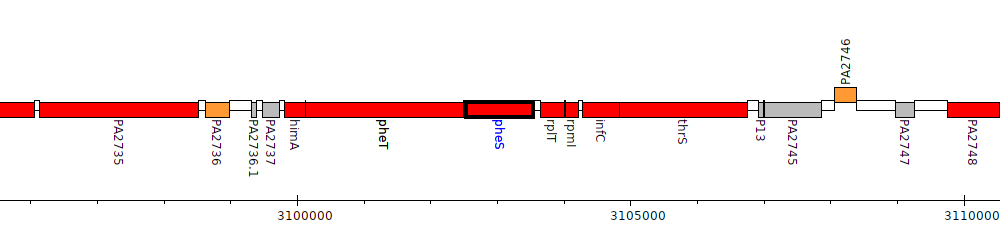
Pseudomonas aeruginosa PAO1, PA2740 (pheS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009094 | L-phenylalanine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0000162 | tryptophan biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006571 | tyrosine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0043039 | tRNA aminoacylation | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004818 | glutamate-tRNA ligase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00468
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006432 | phenylalanyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00468
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000049 | tRNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00468
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0043039 | tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF01409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00468
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004826 | phenylalanine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00468
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Aminoacyl-tRNA biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | TRNA-CHARGING-PWY | tRNA charging | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Phenylalanine, tyrosine and tryptophan biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02912 | Aminoacyl tRNA synthetase class II, N-terminal domain | IPR004188 | Phenylalanine-tRNA ligase, class II, N-terminal | 20 | 87 | 5.3E-27 |
| SUPERFAMILY | SSF46589 | tRNA-binding arm | IPR010978 | Class I and II aminoacyl-tRNA synthetase, tRNA-binding arm | 13 | 99 | 2.51E-23 |
| NCBIfam | TIGR00468 | JCVI: phenylalanine--tRNA ligase subunit alpha | IPR004529 | Phenylalanyl-tRNA synthetase, class IIc, alpha subunit | 38 | 337 | 1.7E-106 |
| PANTHER | PTHR11538 | PHENYLALANYL-TRNA SYNTHETASE | - | - | 89 | 337 | 1.2E-30 |
| SUPERFAMILY | SSF55681 | Class II aaRS and biotin synthetases | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 92 | 338 | 2.68E-82 |
| Hamap | MF_00281 | Phenylalanine--tRNA ligase alpha subunit [pheS]. | IPR022911 | Phenylalanine-tRNA ligase alpha chain 1, bacterial | 8 | 337 | 50.770996 |
| CDD | cd00496 | PheRS_alpha_core | - | - | 107 | 332 | 1.05767E-126 |
| Gene3D | G3DSA:3.30.930.10 | Bira Bifunctional Protein; Domain 2 | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 7 | 338 | 0.0 |
| FunFam | G3DSA:3.30.930.10:FF:000003 | Phenylalanine--tRNA ligase alpha subunit | - | - | 8 | 338 | 0.0 |
| Pfam | PF01409 | tRNA synthetases class II core domain (F) | IPR002319 | Phenylalanyl-tRNA synthetase | 92 | 337 | 1.8E-108 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.