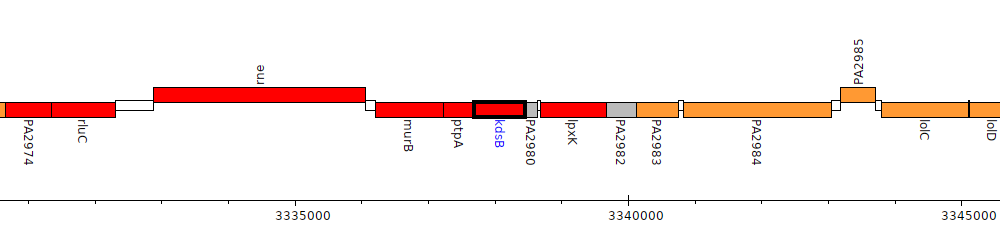
Pseudomonas aeruginosa PAO1, PA2979 (kdsB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003867 | 4-aminobutyrate transaminase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006013 | mannose metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006000 | fructose metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008690 | 3-deoxy-manno-octulosonate cytidylyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00057
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PWY-1269 | CMP-3-deoxy-D-manno-octulosonate biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | KDO-PEP-LIPASYN-PWY | KDO<SUB>2</SUB>-lipid A and peptidoglycan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Fructose and mannose metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00540 | Lipopolysaccharide biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | KDO-NAGLIPASYN-PWY | superpathway of (Kdo)<SUB>2</SUB>-lipid A biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR42866 | 3-DEOXY-MANNO-OCTULOSONATE CYTIDYLYLTRANSFERASE | - | - | 5 | 251 | 3.0E-68 |
| FunFam | G3DSA:3.90.550.10:FF:000011 | 3-deoxy-manno-octulosonate cytidylyltransferase | - | - | 3 | 253 | 4.8E-113 |
| Pfam | PF02348 | Cytidylyltransferase | IPR003329 | Acylneuraminate cytidylyltransferase | 7 | 226 | 1.0E-55 |
| CDD | cd02517 | CMP-KDO-Synthetase | IPR004528 | 3-deoxy-D-manno-octulosonate cytidylyltransferase | 5 | 251 | 1.30278E-130 |
| NCBIfam | TIGR00466 | JCVI: 3-deoxy-manno-octulosonate cytidylyltransferase | IPR004528 | 3-deoxy-D-manno-octulosonate cytidylyltransferase | 4 | 246 | 8.7E-93 |
| SUPERFAMILY | SSF53448 | Nucleotide-diphospho-sugar transferases | IPR029044 | Nucleotide-diphospho-sugar transferases | 5 | 251 | 1.6E-60 |
| Gene3D | G3DSA:3.90.550.10 | Spore Coat Polysaccharide Biosynthesis Protein SpsA; Chain A | IPR029044 | Nucleotide-diphospho-sugar transferases | 1 | 254 | 4.4E-96 |
| Hamap | MF_00057 | 8-amino-3,8-dideoxy-manno-octulosonate cytidylyltransferase [kdsB]. | IPR004528 | 3-deoxy-D-manno-octulosonate cytidylyltransferase | 3 | 253 | 43.99445 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.