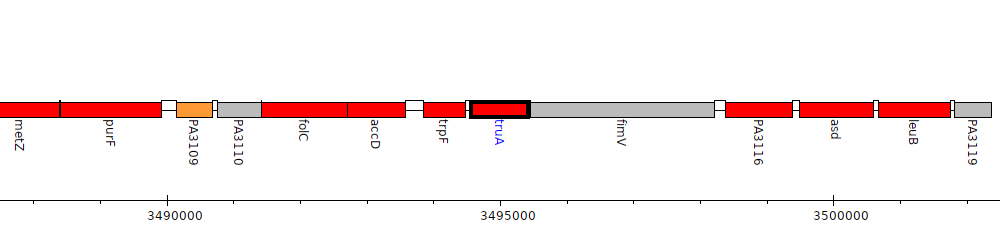
Pseudomonas aeruginosa PAO1, PA3114 (truA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006220 | pyrimidine nucleotide metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0003941 | L-serine ammonia-lyase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd02570
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009451 | RNA modification |
Inferred from Sequence Model
Term mapped from: InterPro:cd02570
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009982 | pseudouridine synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd02570
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0001522 | pseudouridine synthesis |
Inferred from Sequence Model
Term mapped from: InterPro:cd02570
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Pyrimidine metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF001430 | PSU-A | IPR001406 | Pseudouridine synthase I, TruA | 17 | 277 | 6.8E-74 |
| Pfam | PF01416 | tRNA pseudouridine synthase | IPR020097 | Pseudouridine synthase I, TruA, alpha/beta domain | 25 | 120 | 4.1E-5 |
| Hamap | MF_00171 | tRNA pseudouridine synthase A [truA]. | IPR001406 | Pseudouridine synthase I, TruA | 18 | 261 | 34.619537 |
| Gene3D | G3DSA:3.30.70.660 | - | IPR020095 | Pseudouridine synthase I, TruA, C-terminal | 123 | 260 | 9.8E-43 |
| FunFam | G3DSA:3.30.70.580:FF:000001 | tRNA pseudouridine synthase A | - | - | 17 | 122 | 8.0E-41 |
| Pfam | PF01416 | tRNA pseudouridine synthase | IPR020097 | Pseudouridine synthase I, TruA, alpha/beta domain | 160 | 262 | 7.4E-31 |
| Gene3D | G3DSA:3.30.70.580 | - | IPR020094 | Pseudouridine synthase TruA/RsuA/RluB/E/F, N-terminal | 15 | 122 | 2.4E-31 |
| NCBIfam | TIGR00071 | JCVI: tRNA pseudouridine(38-40) synthase TruA | IPR001406 | Pseudouridine synthase I, TruA | 19 | 256 | 5.1E-66 |
| CDD | cd02570 | PseudoU_synth_EcTruA | IPR001406 | Pseudouridine synthase I, TruA | 22 | 261 | 0.0 |
| PANTHER | PTHR11142 | PSEUDOURIDYLATE SYNTHASE | IPR001406 | Pseudouridine synthase I, TruA | 17 | 265 | 7.2E-58 |
| SUPERFAMILY | SSF55120 | Pseudouridine synthase | IPR020103 | Pseudouridine synthase, catalytic domain superfamily | 18 | 275 | 4.32E-91 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.