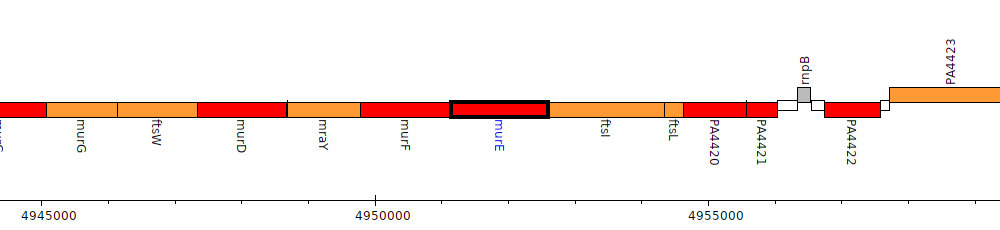
Pseudomonas aeruginosa PAO1, PA4417 (murE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008360 | regulation of cell shape |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53623
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53623
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0051301 | cell division |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016881 | acid-amino acid ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PEP-LIPA-SYN-PWY | peptidoglycan and lipid A precursor biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00550 | Peptidoglycan biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-6387 | UDP-N-acetylmuramoyl-pentapeptide biosynthesis I (meso-DAP-containing) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Peptideglycan biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PEPTIDOGLYCANSYN-PWY | peptidoglycan biosynthesis I (meso-diaminopimelate containing) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | KDO-PEP-LIPASYN-PWY | KDO<SUB>2</SUB>-lipid A and peptidoglycan biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00300 | Lysine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR23135 | MUR LIGASE FAMILY MEMBER | - | - | 3 | 482 | 0.0 |
| Pfam | PF02875 | Mur ligase family, glutamate ligase domain | IPR004101 | Mur ligase, C-terminal | 329 | 412 | 8.9E-23 |
| Gene3D | G3DSA:3.40.1390.10 | - | - | - | 3 | 96 | 1.7E-23 |
| SUPERFAMILY | SSF53623 | MurD-like peptide ligases, catalytic domain | IPR036565 | Mur-like, catalytic domain superfamily | 97 | 326 | 6.41E-77 |
| Gene3D | G3DSA:3.40.1190.10 | - | IPR036565 | Mur-like, catalytic domain superfamily | 97 | 329 | 1.9E-63 |
| Gene3D | G3DSA:3.90.190.20 | - | IPR036615 | Mur ligase, C-terminal domain superfamily | 330 | 483 | 2.1E-57 |
| Hamap | MF_00208 | UDP-N-acetylmuramoyl-L-alanyl-D-glutamate--2,6-diaminopimelate ligase [murE]. | IPR005761 | UDP-N-acetylmuramoylalanyl-D-glutamate-2,6-diaminopimelate ligase | 3 | 479 | 35.0854 |
| NCBIfam | TIGR01085 | JCVI: UDP-N-acetylmuramyl-tripeptide synthetase | IPR005761 | UDP-N-acetylmuramoylalanyl-D-glutamate-2,6-diaminopimelate ligase | 17 | 479 | 0.0 |
| Pfam | PF08245 | Mur ligase middle domain | IPR013221 | Mur ligase, central | 106 | 306 | 2.6E-53 |
| Pfam | PF01225 | Mur ligase family, catalytic domain | IPR000713 | Mur ligase, N-terminal catalytic domain | 18 | 94 | 1.7E-10 |
| SUPERFAMILY | SSF63418 | MurE/MurF N-terminal domain | IPR035911 | MurE/MurF, N-terminal | 11 | 95 | 6.28E-21 |
| SUPERFAMILY | SSF53244 | MurD-like peptide ligases, peptide-binding domain | IPR036615 | Mur ligase, C-terminal domain superfamily | 329 | 480 | 1.45E-47 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.