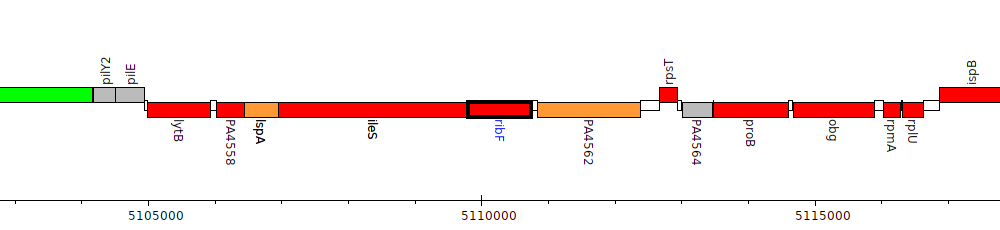
Pseudomonas aeruginosa PAO1, PA4561 (ribF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008531 | riboflavin kinase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
9473052 | Reviewed by curator |
| Molecular Function | GO:0003919 | FMN adenylyltransferase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
9473052 | Reviewed by curator |
| Molecular Function | GO:0003919 | FMN adenylyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF06574
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009231 | riboflavin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF06574
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008531 | riboflavin kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF82114
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | RIBOSYN2-PWY | flavin biosynthesis I (bacteria and plants) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00740 | Riboflavin metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Riboflavin metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY-5523 | 5,6-dimethylbenzimidazole biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P381-PWY | adenosylcobalamin biosynthesis II (late cobalt incorporation) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR22749 | RIBOFLAVIN KINASE/FMN ADENYLYLTRANSFERASE | IPR023468 | Riboflavin kinase | 7 | 304 | 1.3E-48 |
| Gene3D | G3DSA:2.40.30.30 | - | IPR023465 | Riboflavin kinase domain superfamily | 186 | 309 | 2.2E-37 |
| SUPERFAMILY | SSF52374 | Nucleotidylyl transferase | - | - | 17 | 178 | 1.69E-31 |
| Pfam | PF06574 | FAD synthetase | IPR015864 | FAD synthetase | 15 | 167 | 1.0E-54 |
| CDD | cd02064 | FAD_synthetase_N | IPR015864 | FAD synthetase | 17 | 197 | 1.47763E-85 |
| SMART | SM00904 | Flavokinase_2 | IPR015865 | Riboflavin kinase domain, bacterial/eukaryotic | 183 | 307 | 8.6E-60 |
| NCBIfam | TIGR00083 | JCVI: riboflavin biosynthesis protein RibF | IPR002606 | Riboflavin kinase, bacterial | 20 | 305 | 1.2E-96 |
| PIRSF | PIRSF004491 | RibF_RibC | IPR002606 | Riboflavin kinase, bacterial | 1 | 310 | 1.2E-103 |
| FunFam | G3DSA:3.40.50.620:FF:000021 | Riboflavin biosynthesis protein | - | - | 1 | 185 | 6.1E-66 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 1 | 185 | 1.1E-62 |
| SUPERFAMILY | SSF82114 | Riboflavin kinase-like | IPR023465 | Riboflavin kinase domain superfamily | 174 | 306 | 1.46E-43 |
| Pfam | PF01687 | Riboflavin kinase | IPR015865 | Riboflavin kinase domain, bacterial/eukaryotic | 184 | 305 | 9.3E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.