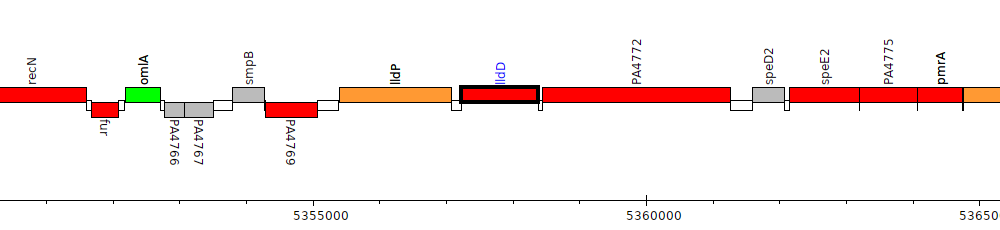
Pseudomonas aeruginosa PAO1, PA4771 (lldD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004459 | L-lactate dehydrogenase activity |
Inferred from Genetic Interaction
Term mapped from: PseudoCAP:PA2382
|
ECO:0000316 genetic interaction evidence used in manual assertion |
30066495 | Reviewed by curator |
| Biological Process | GO:0019516 | lactate oxidation | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8407843 | Reviewed by curator |
| Molecular Function | GO:0004459 | L-lactate dehydrogenase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
18473 | Reviewed by curator |
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01559
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006089 | lactate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01559
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000138
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004457 | lactate dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01559
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | LCTACACAT-PWY | lactate oxidation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P122-PWY | heterolactic fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P461-PWY | hexitol fermentation to lactate, formate, ethanol and acetate | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Pyruvate metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-5481 | pyruvate fermentation to lactate | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00620 | Pyruvate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P41-PWY | pyruvate fermentation to acetate and lactate I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd02809 | alpha_hydroxyacid_oxid_FMN | IPR012133 | Alpha-hydroxy acid dehydrogenase, FMN-dependent | 8 | 371 | 0.0 |
| PIRSF | PIRSF000138 | Al-hdrx_acd_dh | IPR012133 | Alpha-hydroxy acid dehydrogenase, FMN-dependent | 2 | 379 | 2.2E-122 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 3 | 379 | 7.6E-124 |
| SUPERFAMILY | SSF51395 | FMN-linked oxidoreductases | - | - | 7 | 374 | 2.56E-98 |
| Pfam | PF01070 | FMN-dependent dehydrogenase | IPR000262 | FMN-dependent dehydrogenase | 13 | 375 | 1.8E-125 |
| PANTHER | PTHR10578 | S -2-HYDROXY-ACID OXIDASE-RELATED | - | - | 5 | 376 | 7.7E-104 |
| Hamap | MF_01559 | L-lactate dehydrogenase [lldD]. | IPR020920 | L-lactate dehydrogenase, bacterial | 1 | 380 | 75.02739 |
| NCBIfam | NF033901 | NCBIFAM: FMN-dependent L-lactate dehydrogenase LldD | IPR020920 | L-lactate dehydrogenase, bacterial | 1 | 377 | 0.0 |
| FunFam | G3DSA:3.20.20.70:FF:000029 | L-lactate dehydrogenase | - | - | 2 | 380 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.