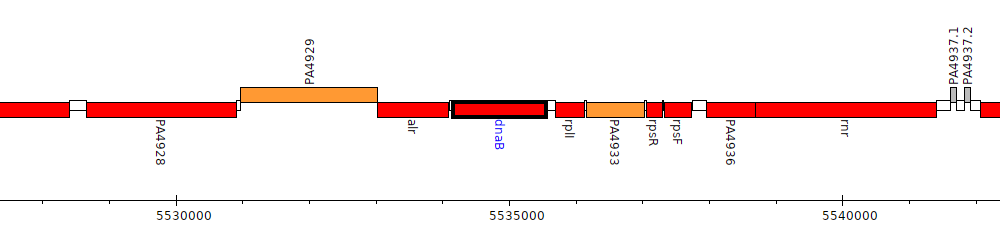
Pseudomonas aeruginosa PAO1, PA4931 (dnaB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006270 | DNA replication initiation | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
345277 | Reviewed by curator |
| Biological Process | GO:0006270 | DNA replication initiation | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
345277 | Reviewed by curator |
| Biological Process | GO:0006260 | DNA replication | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3007474 | Reviewed by curator |
| Molecular Function | GO:0003678 | DNA helicase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3007474 | Reviewed by curator |
| Molecular Function | GO:0016887 | ATPase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11061975 | Reviewed by curator |
| Molecular Function | GO:0005524 | ATP binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
11061975 | Reviewed by curator |
| Biological Process | GO:0032508 | DNA duplex unwinding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3007474 | Reviewed by curator |
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PF03796
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003678 | DNA helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF03796
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF03796
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00665
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae03030 | DNA replication | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48024 | N-terminal domain of DnaB helicase | IPR036185 | DNA helicase, DnaB-like, N-terminal domain superfamily | 18 | 129 | 1.31E-40 |
| FunFam | G3DSA:3.40.50.300:FF:000076 | Replicative DNA helicase | - | - | 177 | 459 | 0.0 |
| CDD | cd00984 | DnaB_C | IPR007694 | DNA helicase, DnaB-like, C-terminal | 197 | 453 | 0.0 |
| NCBIfam | TIGR00665 | JCVI: replicative DNA helicase | IPR007692 | DNA helicase, DnaB type | 20 | 455 | 0.0 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 196 | 446 | 7.67E-53 |
| Pfam | PF03796 | DnaB-like helicase C terminal domain | IPR007694 | DNA helicase, DnaB-like, C-terminal | 197 | 453 | 1.7E-113 |
| Gene3D | G3DSA:1.10.860.10 | DNAb Helicase; Chain A | IPR016136 | DNA helicase DnaB, N-terminal/DNA primase DnaG, C-terminal | 13 | 178 | 1.9E-51 |
| FunFam | G3DSA:1.10.860.10:FF:000001 | Replicative DNA helicase | - | - | 13 | 178 | 9.2E-57 |
| Pfam | PF00772 | DnaB-like helicase N terminal domain | IPR007693 | DNA helicase, DnaB-like, N-terminal | 20 | 121 | 1.7E-37 |
| PANTHER | PTHR30153 | REPLICATIVE DNA HELICASE DNAB | - | - | 14 | 457 | 0.0 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 179 | 459 | 1.4E-107 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.